|
|
|
|
Our on-going research concerns the reconstruction and analysis of biological and clinical networks. We develop information-theoretic methods and machine learning tools to infer interpretable causal graphical models from large scale genomic data (single cell multi-omic data), live-cell imaging data (tumor-on-chip experiments) as well as medical records of patients. |
 |
|||
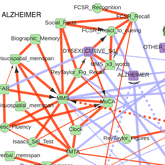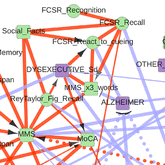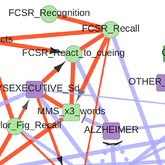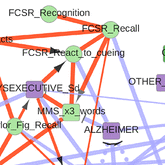 |
Information maximization in mixed-type data from medical records |
||
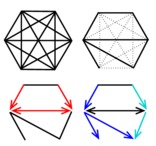 |
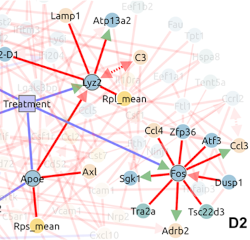
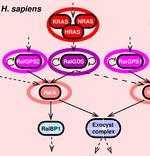
Robust reconstruction of causal, non-causal or mixed networks |
||
 |
|||
 |
|||
 |
|||
 |
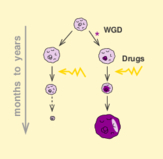
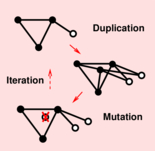
Identification of ohnologs from whole genome duplication |
||
 |
|||
 |
|||
|
|
|||
 |
|||
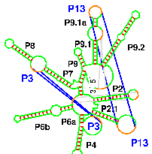 |
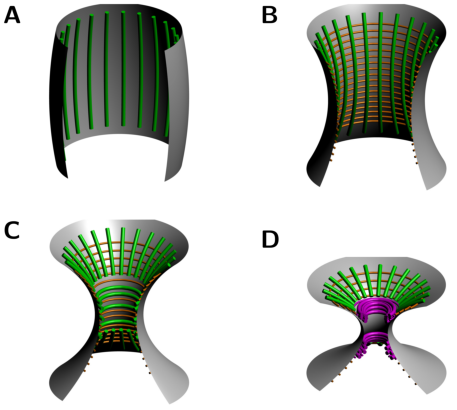
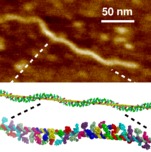
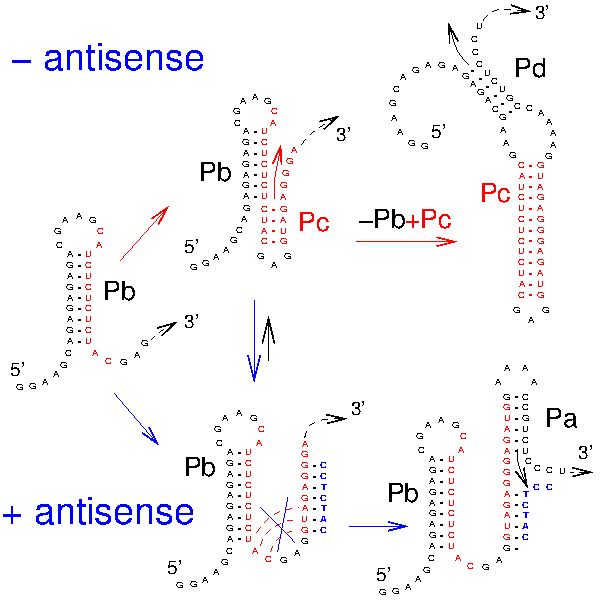
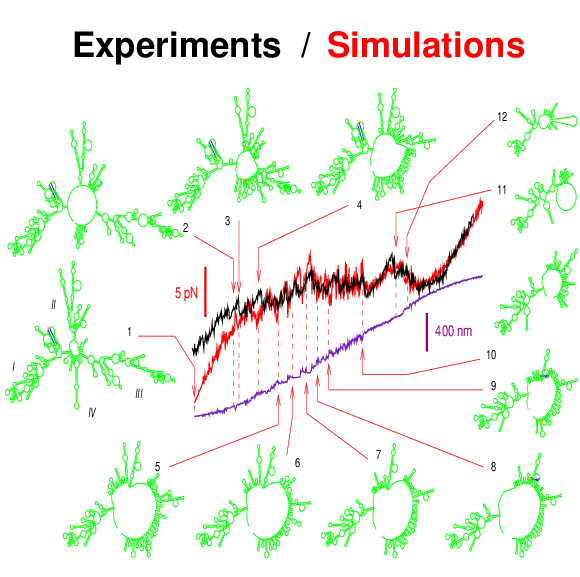
RNA folding including pseudoknots and "real knots" |
||
 |
|||
 |
|||
 |
|||
 |

