|
|
|
|
More reliable and interpretable causal discovery methods The reliability and interpretability of machine learning methods have become major issues in Artificial Intelligence. These questions are particularly important for AI approaches applied to sensitive data, such as medical data of patients for which AI-assisted recommendations can hardly rely on black box classifiers only and need to be explainable in terms of intelligible medical rationales.
Our specific interest lies in the reliability and interpretability of causal discovery methods from purely observational data.
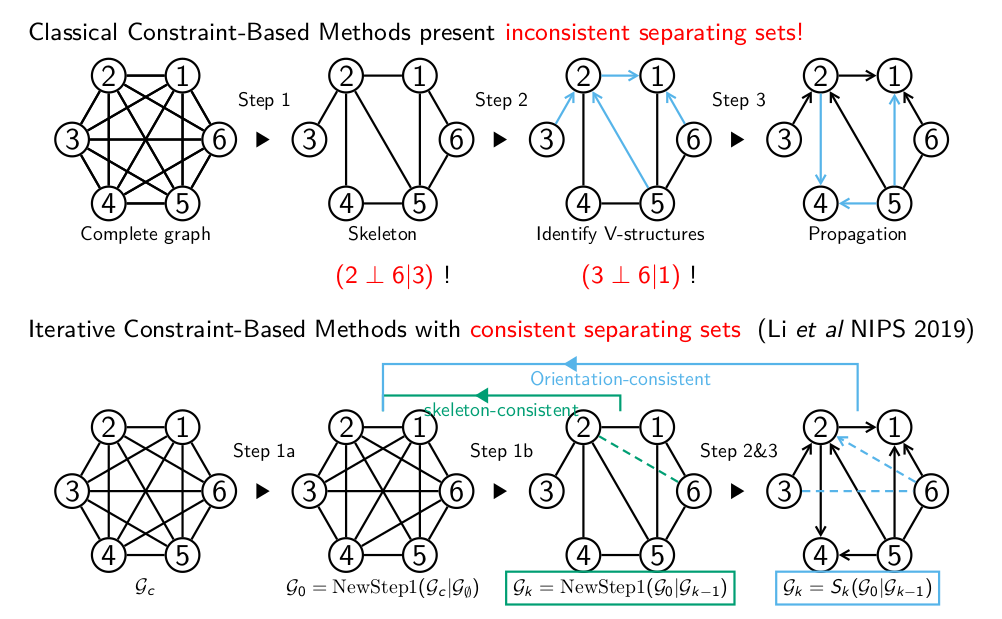
These constraint-based structure learning methods start with a first step, which consists in pruning a complete graph to obtain an undirected graph skeleton, that is subsequently oriented, Figure 1. All causal discovery methods perform this first step of removing dispensable edges, iteratively, whenever a separating set and corresponding conditional independence can be found. Yet, causal discovery methods lack robustness over sampling noise and are prone to uncover spurious conditional independences in finite datasets. In particular, there is no guarantee that the separating sets identified during the iterative pruning step remain consistent with the final graph in terms of indirect paths through these nodes.
| ||
|
|
Figure 1. Iterative constraint-based structure learning algorithms with consistent separating sets. (Li et al. NeurIPS 2019) |
||
|
Ensuring the consistency of separating sets can be done at a limited complexity cost, through the use of block-cut tree decomposition of graph skeletons, and is found to increase their validity in terms of actual d-separation. It also significantly improves the sensitivity of constraint-based methods while retaining good overall structure learning performance. Finally and foremost, ensuring sepset consistency improves the interpretability of constraint-based models for real-life applications, as discussed below.
|
|||
|
|
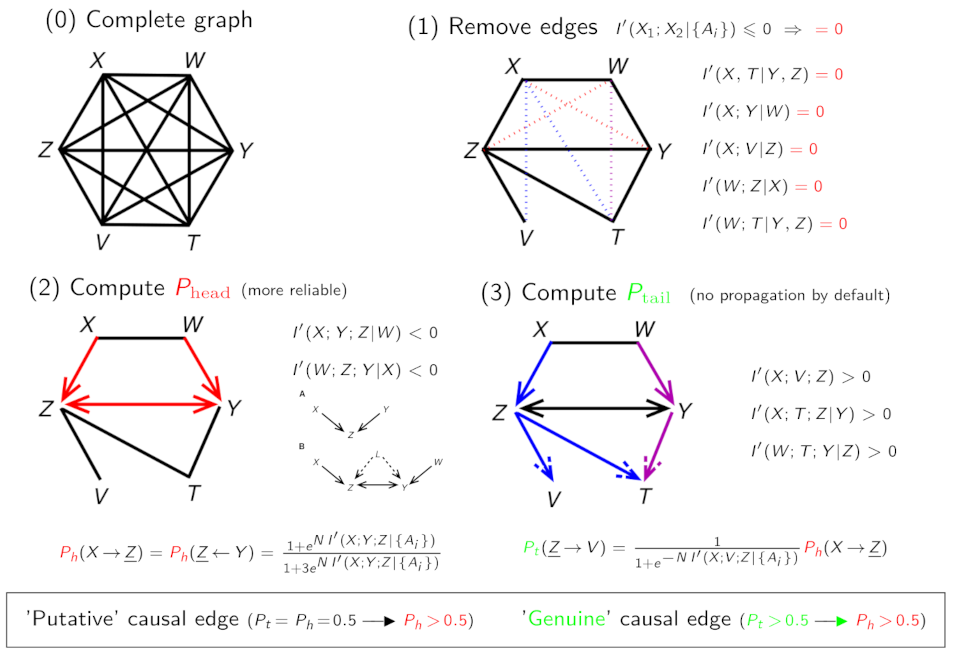
Figure 2. Algorithm scheme of iMIIC. (Ribeiro-Dantas et al. 2024). |
||
|
First, iMIIC quantitatively improves the reliability of inferred orientations, based on a general information-theoretic principle (Cabeli et al. 2021, Ribeiro-Dantas et al. 2024). This implies to rectify all negative mutual information regularized for finite size effect, in Figure 2 (step 1), and results in only a few percents of false positive orientations, in Figure 2 (step 2), on challenging benchmarks adapted from real-world healthcare data. Second, iMIIC is uniquely able to distinguish "genuine" causes from "putative" and "latent" causal effects, in Figure 2 (step 3). This is an essential distinction to disambiguate the causal interpretation of oriented edges in inferred networks. Third, iMIIC quantifies indirect effects, while ensuring their consistency with the global network structure through the iterative scheme outlined in Figure 1. This is important to interpret indirect contributions in term of indirect paths through the corresponding contributor nodes in the inferred network, which is generally not possible with other causal discovery methods.
|
|||
|
|
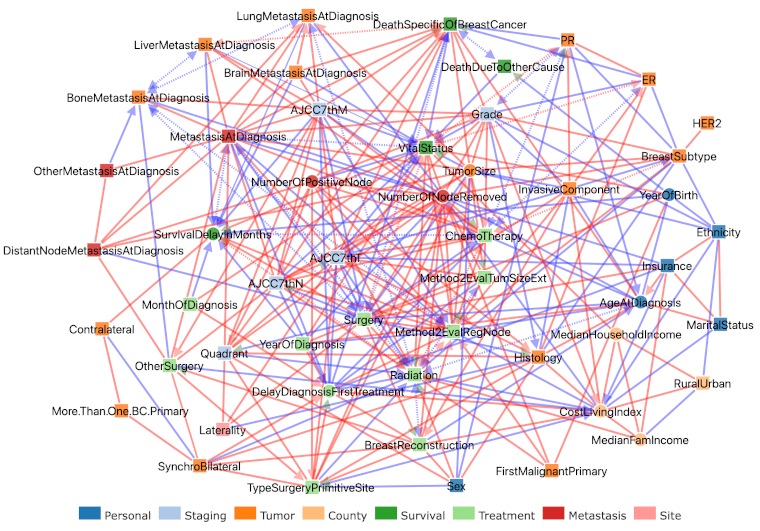
Figure 3. SEER breast cancer networks inferred by iMIIC. The 51 node network inferred by iMIIC from SEER dataset including 396,179 breast cancer patients diagnosed between 2010 and 2016. An expert knowledge validation of the causal effects inferred by iMIIC indicates that about 90% of predicted causal effects are correct, while an additional 8% of cause-effect relations seem plausible, based on clinical and epidemiological knowledge (Ribeiro-Dantas et al. 2024). |
||
|
Hence, iMIIC provides a detailed and validated interpretation across all variables selected in this nation-wide cohort of 400,000 breast cancer patients. This exhaustive analysis uncovers many expected causal relations, such as the adverse consequence of metastasis and the protecting effect of ER+ and specifically PR+ status on death due to breast cancer, or the fact that year of birth is the primary reason for death due to other causes by the end of the study.
|
|||
|
|
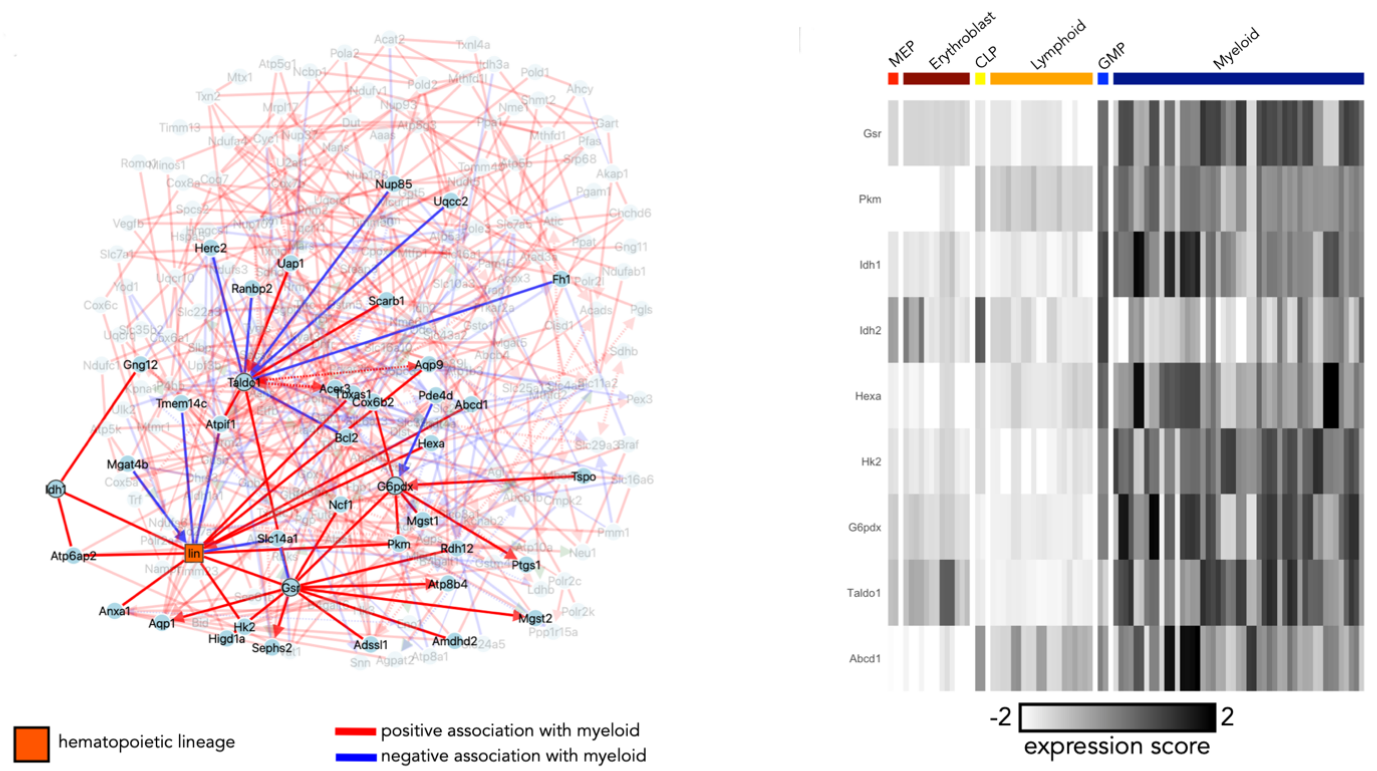
Figure 4. On the role of metabolism in hematopoietic differentiation. iMIIC network uncovers metabolic gene association with myeloid / erythroid-lymphoid differentiation fate (left), using bulk RNA seq profiles from the haemopedia database. This suggests that metabolic heterogeneity is an important regulatory component of hematopoiesis, which was subsequently demonstrated in vitro and in vivo (right) (Cosgrove et al. bioRxiv 2023). |
||
|
Another recent application concerns the analysis of scRNAseq data from a cellular therapy against multiple sclerosis in a mouse model in collaboration with the Fillatreau lab, Institut Necker Enfants Malades (Manfroi et al. 2026). Other recent scRNAseq applications concern the analysis of gene regulatory pathways driving Extracellular Matrix remodeling by cancer cells and fibroblasts under macrophage depletion (Fusilier et al. 2026), and an integration of scRNAseq data with image-derived synchronicity markers to investigate cellular memory in human hematopoietic stem cells (Donada et al. 2026). Finally, we have also developed the CausalCCC web server to reconstruct gene-gene interaction pathways across two or more interacting cell types from single-cell or spatial transcriptomic data, Figure 5 (Dupuis & Debeaupuis et al. 2025).
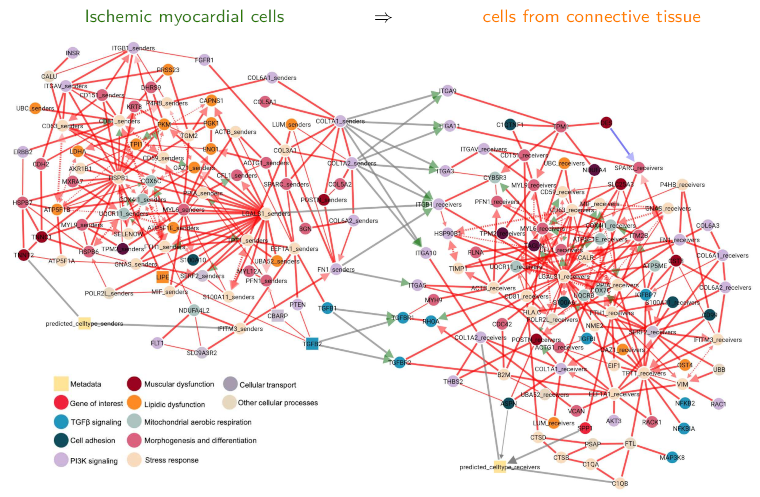
Figure 5. CausalCCC analysis: CausalCCC network reconstruction in Myocardiac Infraction, using (Kuppe et al. 2022) spatial transcriptomic data (10X visum, Patient 9). See CausalCCC web server (Dupuis & Debeaupuis et al. 2025). Extension to time series data with applications to time-lapse microscopy imaging data Another recent extension of MIIC algorithm concerns its temporal version (tMIIC) to analyze time series data, in either stationary (Simon et al. 2025) or non-stationary (Parent et al. 2025) regimes. We have applied this temporal causal discovery approach to time-lapse microscopy images of tumor-on-chip data (Simon et al. 2025) and tumoroid data (Parent et al. 2025). In particular, we have developed the CausalXtract pipeline to reconstruct causal networks from images of individual cancer cells in order to understand how treatment and the tumor microenvironment affect their morphology, division and death, as well as their interactions with immune cells, Figure 6 (Simon et al. 2025). We also reconstructed non-stationary temporal networks from bright field images of lab-grown tumor spheroids (tumoroids) in order to study their response to drug treatment over time and identify the morphological features that are most informative of drug efficacy (Parent et al. 2025). |
|||
|
|
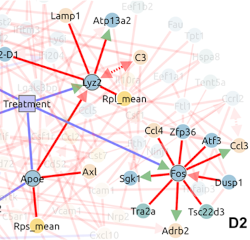
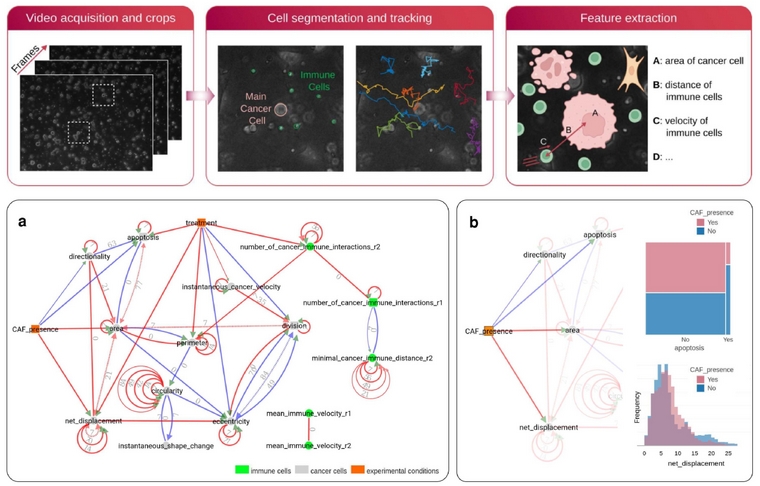
Figure 6. CausalXtract pipeline (Simon et al. 2025) first segments individual cancer and immune cells and extract morphodynamic features from time-lapse images. a. A temporal MIIC network is then reconstructed from the extracted features and the experimentally controlled variables, such as Treatment and the presence Cancer Associated Fibroblasts (CAF). b. In particular, we found that CAFs in the tumor microenvironment directly inhibit cancer cell apoptosis, independently from treatment, which had not been reported so far (Simon et al. 2025). |
||
|
Related Publications |
|