|
|
|
|
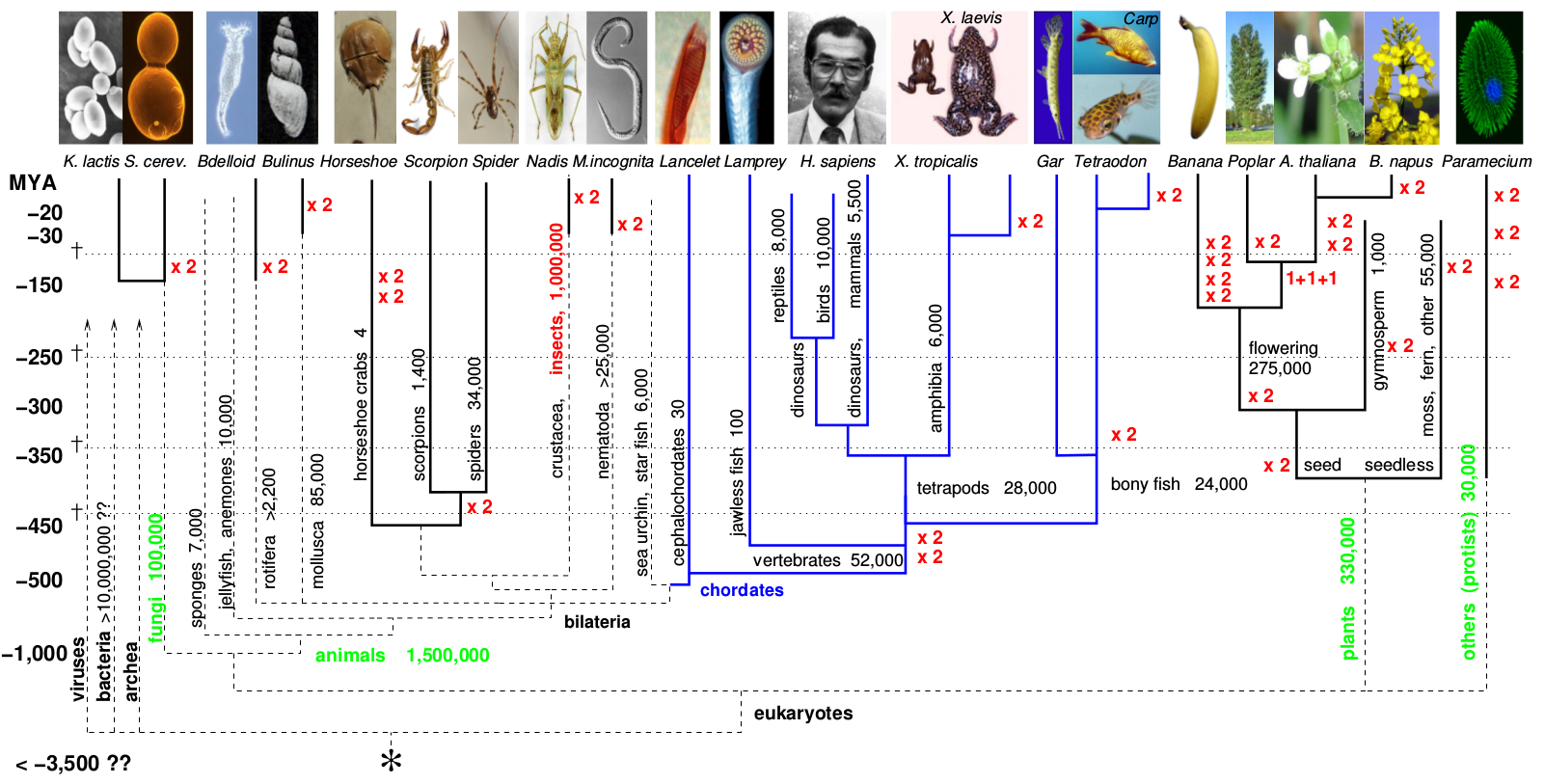
Identification of ohnologs from whole genome duplication Whole genome duplications (WGD) have now been firmly established in all major eukaryotic kingdoms. In particular, all vertebrates descend from two rounds of WGDs, that occurred in their jawless ancestor some 500 MY ago, Figure 1. |
||
|
Figure 1. Whole genome duplications in evolution. Whole genome duplications have occurred repeatedly in the course of eukaryote evolution (Singh et al PLoS Comput Biol 2015). |
|||
|
|
|||
|
|
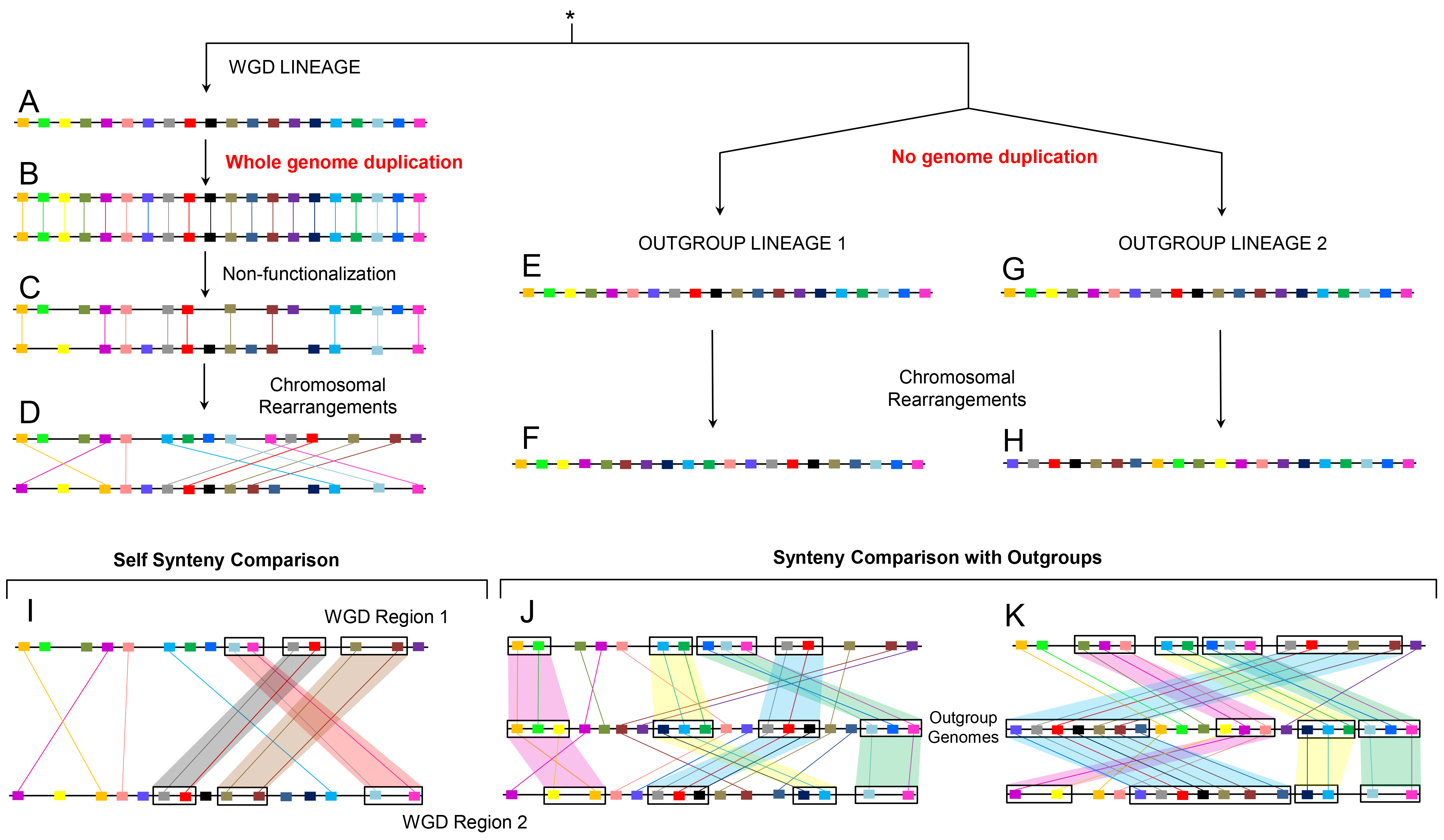
Figure 2. Evolution after WGD and identification of ohnologs. Evolution after WGD and identification of ohnologs using content-based synteny comparison. The genomes of three lineages sharing a common ancestor are shown. Orthologs and paralogs have been depicted by the same color. The WGD lineage (A) underwent whole genome duplication (B) followed by non-functionalization (C) and genome rearrangements (D) leading to the current intragenomic content-based synteny (I). By contrast, the two outgroup genomes without WGD (E, G) experienced lineage specific genome rearrangements (F, H) leading to 1-to-2 content-based synteny pattern with the WGD lineage (J, K). Note, that some ohnolog pairs (D) are only identified by one of the two outgroups (J or K) due to lineage specific rearrangements (Singh et al. PLoS Comput Biol 2015). |
||
|
|
|||
|
|
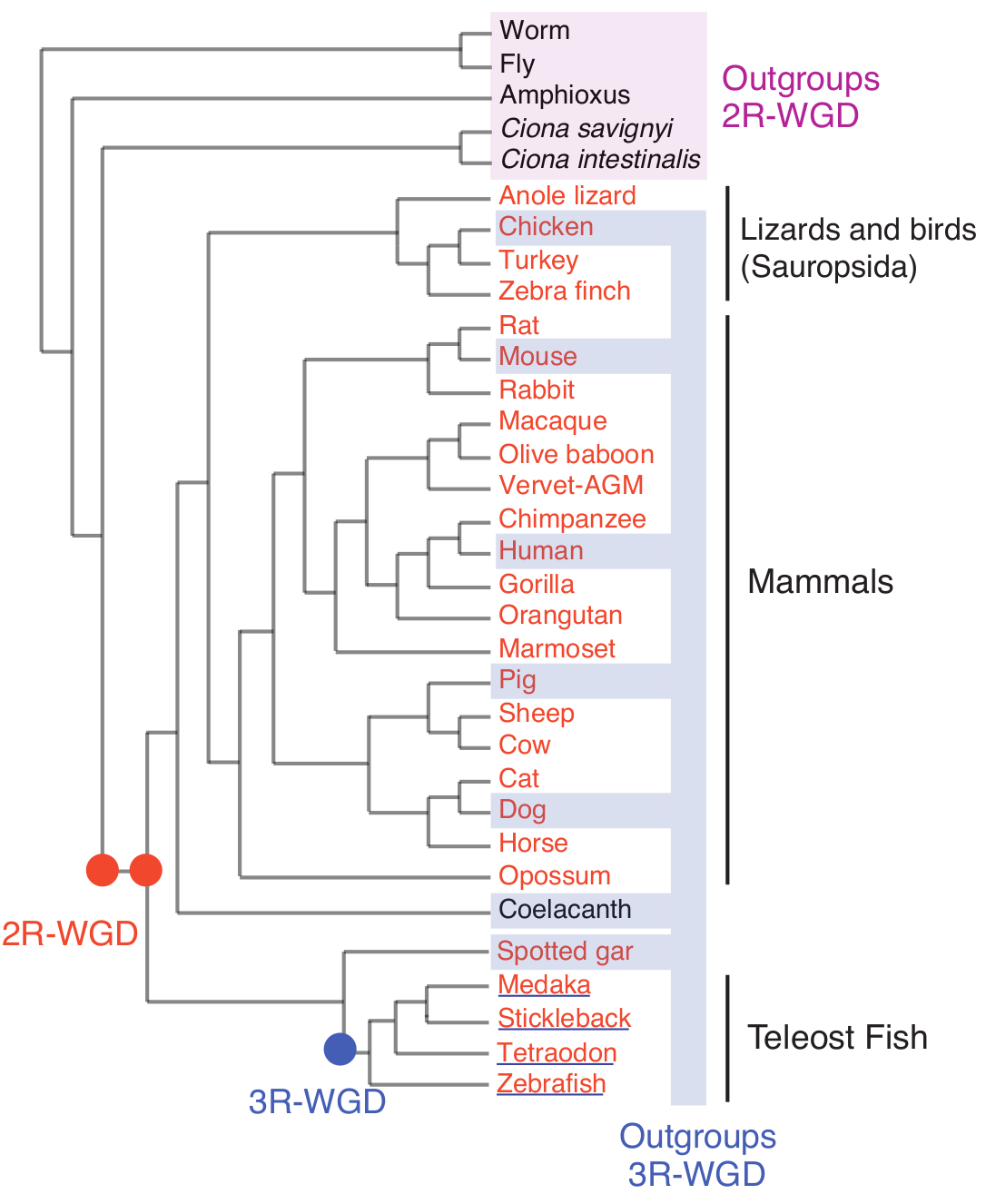
Figure 3. Schematic tree for the 27 vertebrates including 4 teleost fish and 5 invertebrate outgroup organisms. Vertebrates analysed for 2R-WGD are in orange, and teleost fish species analysed for 3R-WGD are underlined. Outgroup species used to identify 2R- and 3R-ohnologs have been highlighted. (Singh & Isambert, Nucleic Acid Res 2020). |
||
|
|
|||
|
|
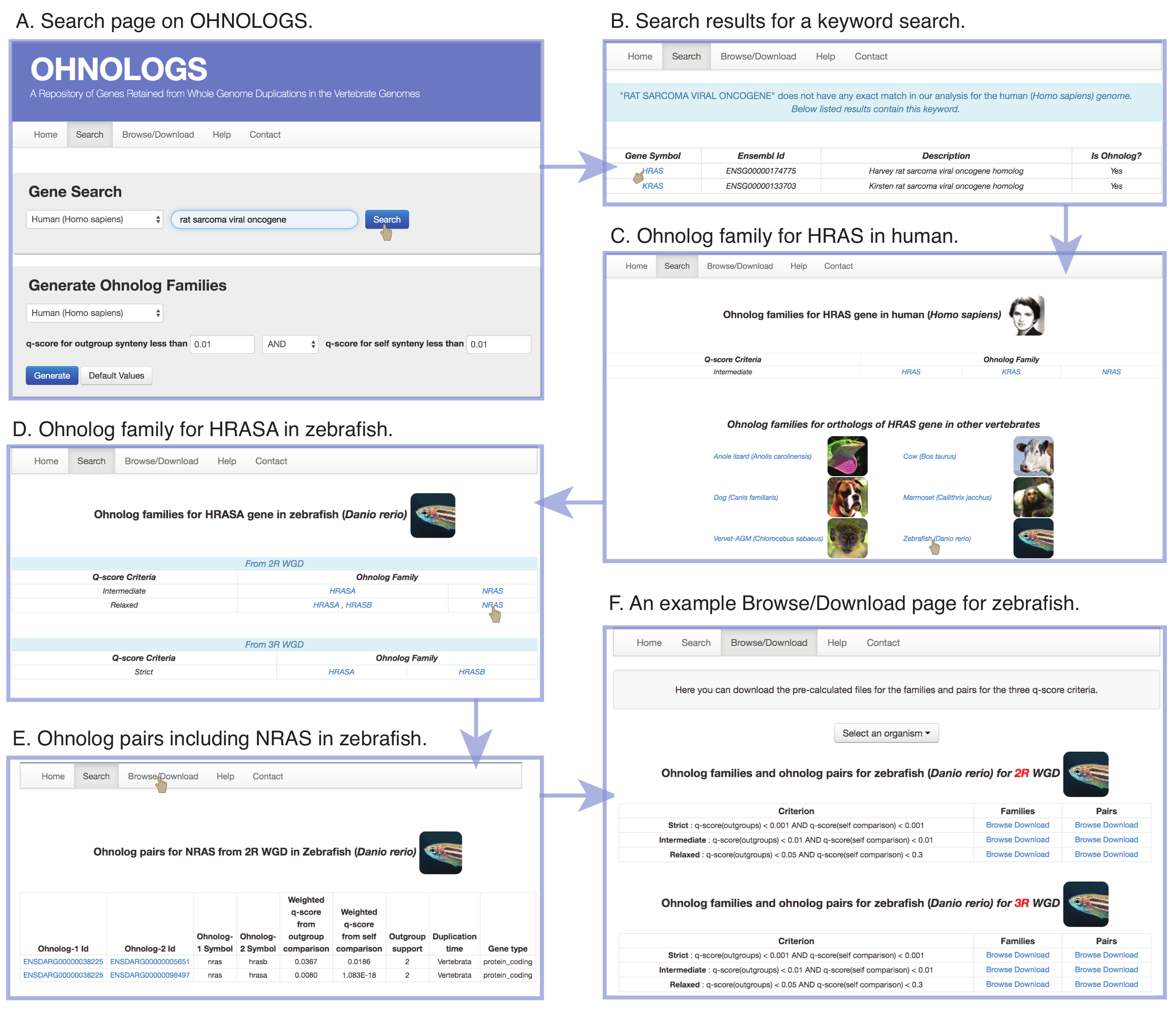
Figure 4. Navigating the OHNOLOGS database. (A) Screenshot of the search page. (B) Result page for a keyword search of 'rat sarcoma viral oncogene' shows the matching genes in human. (C) Ohnolog family page for HRAS gene in the human genome. (D) From the family page, users can navigate to ortholog families in other vertebrates, e.g. zebrafish HRASA. (E) Ohnolog pair page for zebrafish for NRAS gene. (F) Browse/Download page for zebrafish showing both 2R and 3R-ohnolog pairs and families for all the three criteria (Singh & Isambert, Nucleic Acid Res 2020). |
||
|
|
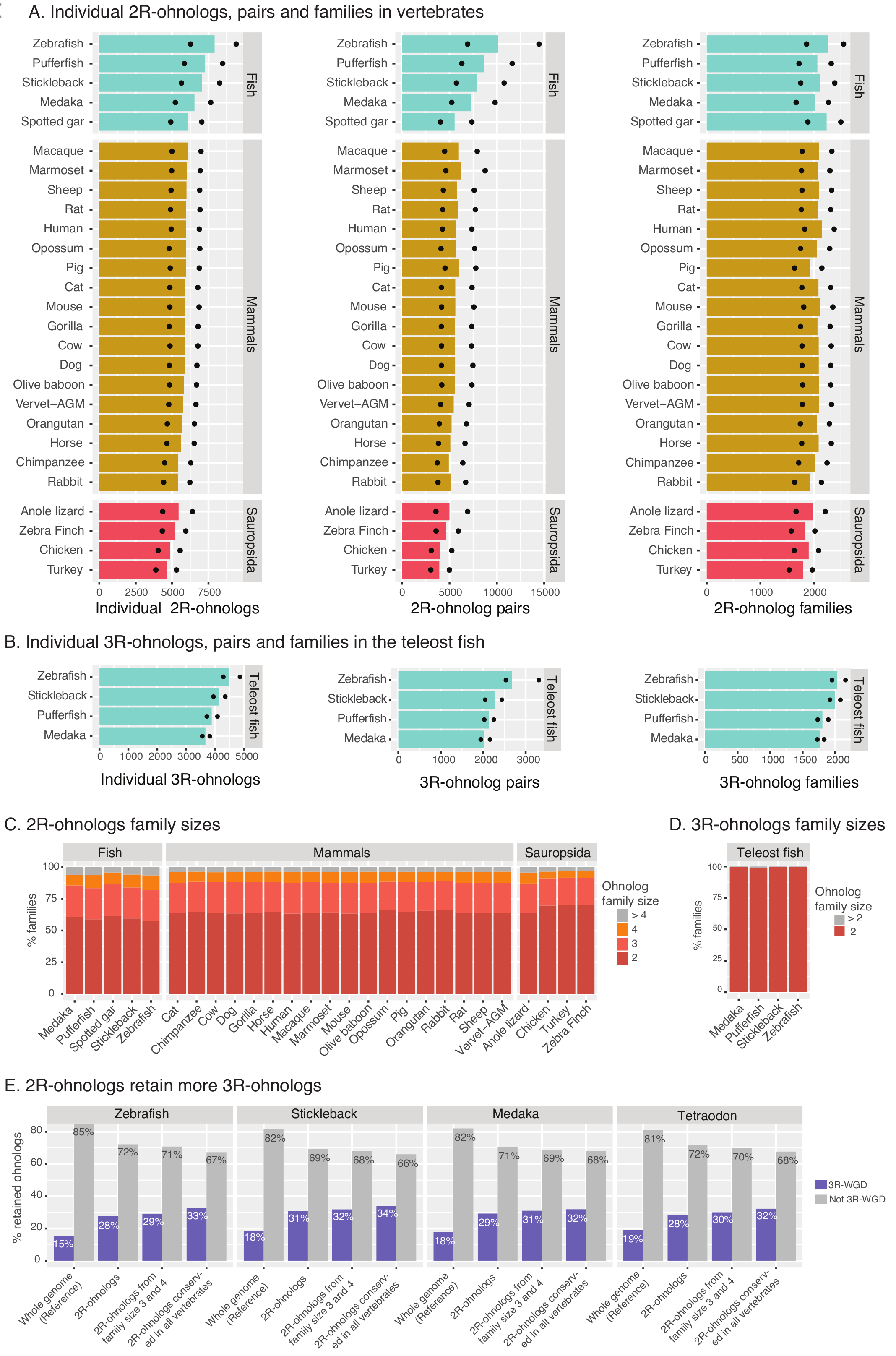
Figure 5. Description of the ohnolog genes, pairs and families in the database. (A) Number of retained individual 2R-ohnolog genes, pairs and families in all the 27 vertebrates. Bars represent the numbers from the intermediate criterion. Ohnologs from strict and relaxed criteria are indicated by dots. (B) Number of retained individual 3R-ohnolog genes, pairs and families in the four teleost fish species. Bars represent the numbers from the intermediate criterion. Ohnologs from strict and relaxed criteria are indicated by dots. (C) Size of the 2R-ohnolog families from the intermediate criterion in vertebrates. Note that a vast majority of the families are of size 2, 3 or 4. (D) Sizes of the 3R-ohnolog families from the intermediate criterion in the teleost fish hardly exceed size two. (E) The 2R-ohnologs are significantly more likely to retain 3R-ohnologs, compared to genome-average. The retention of 3R-ohnologs is even higher for the 2R-ohnologs that belong to family size 3 or 4, and for 2R-ohnologs conserved in all the 27 vertebrates. All the P-values are <1e-41, Chi-square test. Family counts are from the intermediate criterion (Singh & Isambert, Nucleic Acid Res 2020). |
||
|
Related Publications |
|