|
|
|
|
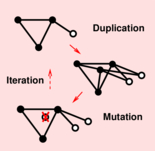
Evolution of large biomolecular networks We investigated the properties of large biomolecular networks and their evolution due to single gene and whole genome duplications, that occurred repeatedly in the course of eukaryote evolution. Our early theoretical analyses focussed on duplication-divergence models to account for the generic properties of biomolecular networks, Figure 1. We have shown that duplication-divergence processes bring not only genetic novelty but also evolutionary constraints that restrict by construction the emerging properties of biomolecular networks. In particular, we demonstrated that networks with evolutionary conserved genes display also necessary topological properties by construction (such as hubs and scale-free degree distribution), Evlampiev et al PNAS 2008; Stein et al PRE 2011; Evlampiev et al BMC Syst Biol 2007. We are also interested in the evolution of transcription networks and study the regulatory conflicts that arise through duplication of transcription factors and autoregulators, Cosentino-Lagomarsino et al PNAS 2007. |
||
|
|
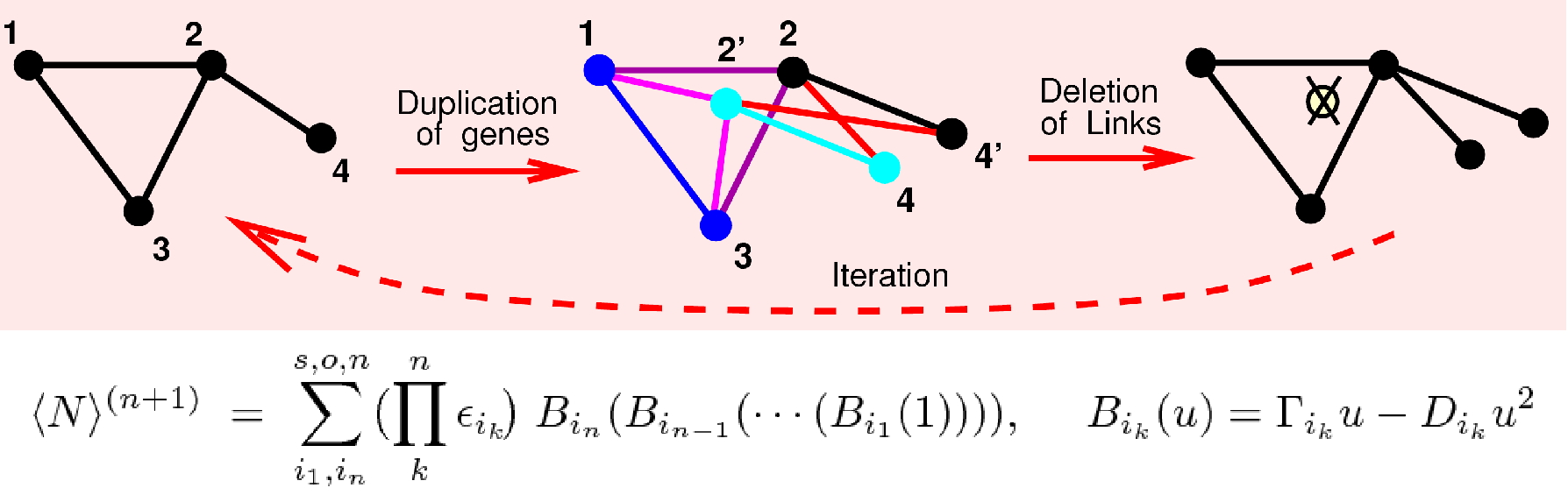
Figure 1. Model of biomolecular networks under duplication-divergence evolution. Biomolecular network evolution by gene and genome duplication can be formally shown to be equivalent to a summation over random compositions of logistic maps, which can be solved analytically in the asymptotic limit of large networks (Evlampiev et al. PNAS 2008; Stein et al. PRE 2011). |
||
|
|
|||
|
Related Publications |
|