| |
|
|
Ancient genome duplications and dominant genetic diseases
In collaboration with the Camonis lab, we analysed the evolutionary constraints on the Ras-Ral signaling networks implicated in cancer, Figure 1. We investigated the evidence that the emerging properties of these signaling pathways might actually reflect their susceptibility to oncogenic mutations and thus their implication in cancer.
|
|
|
|
|
| |
|
|
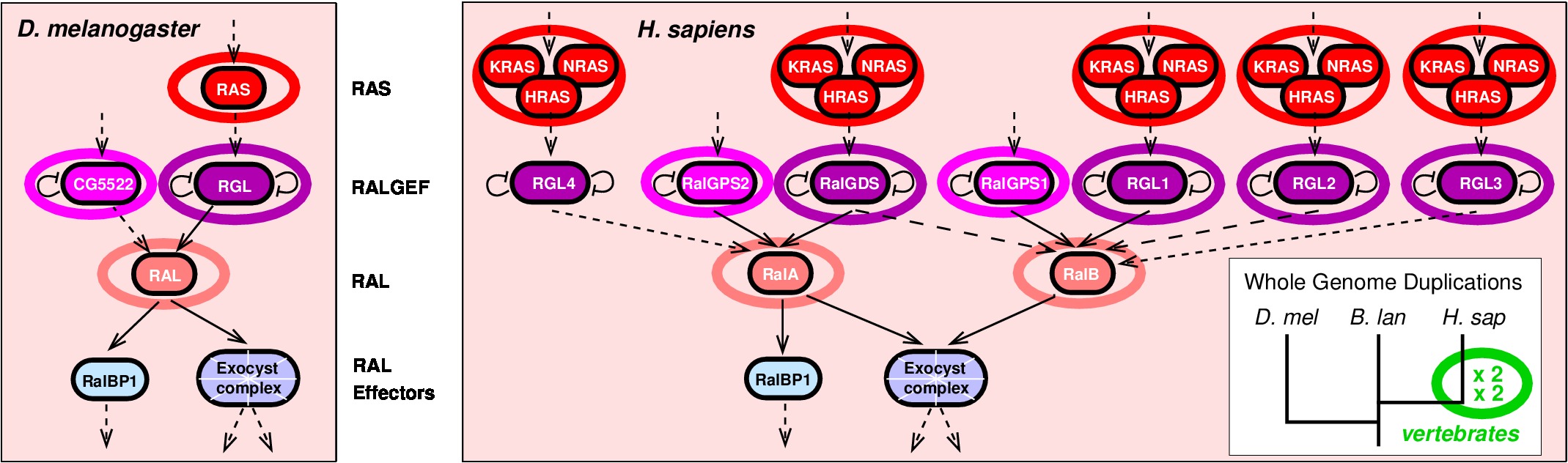
Figure 1.
Expansion of signaling networks by whole genome duplication.
Ras-Ral pathways expansion in vertebrates occurred almost exclusively through two rounds of whole genome duplication in early vertebrates. Adapted from Affeldt et al. Med Sci 2013.
|
| |
|
|
|
|
|
|
|
| |
|
|
We found, in particular, that "dangerous" gene families, defined as prone to dominant deleterious mutations (as the Ral-Ras pathway genes in Figure 1), have been greatly expanded through two rounds of whole genome duplication (WGD) in early vertebrates, Figure 2. By constrast, no such bias is observed for recessive disease genes, Figure 2C.
|
|
|
|
|
| |
|
|
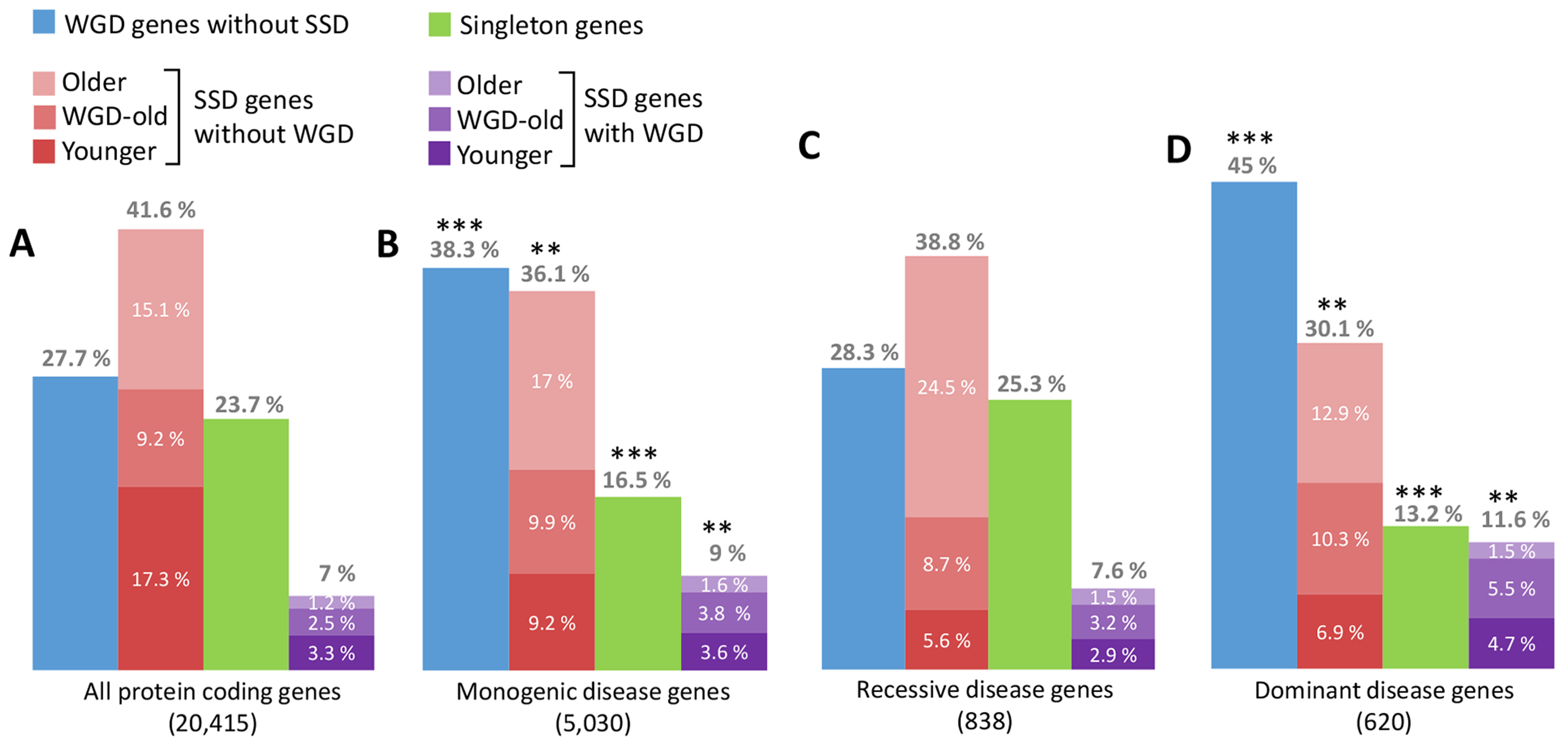
Figure 2.
Expansion of gene families prone to dominant diseases through whole genome duplication (WDG) .
Gene families prone to dominant diseases have been greatly expanded through two rounds of whole genome duplications (WGD, blue), that occurred at the onset of vertebrates. By contrast, genes with recent small scale duplicates (SSD, dark red) are less associated to genetic diseases presumably due to functional backup from recent SSD. Note that few genes (7%) have retained both WDG and SSD duplicates (violet), hinting at different (and mutually exclusive) evolutionary selection scenarios for the retention of WDG and SSD duplicates, Figure 4 (Singh et al. PLoS Comput Biol 2014, Singh et al. Cell Rep 2012).
|
| |
|
|
|
|
|
|
|
| |
|
|
An earlier alternative hypothesis to account for the retention of ohnolog genes from WGD is related to the dosage balance constraint experienced by proteins participating to functional complexes, Figure 3.
However, quantitative Mediation Analysis demonstrates that the retention of many ohnologs suspected to be dosage balanced is in fact indirectly mediated by their susceptibility to deleterious mutations, Figure 3.
|
|
|
|
|
| |
|
|
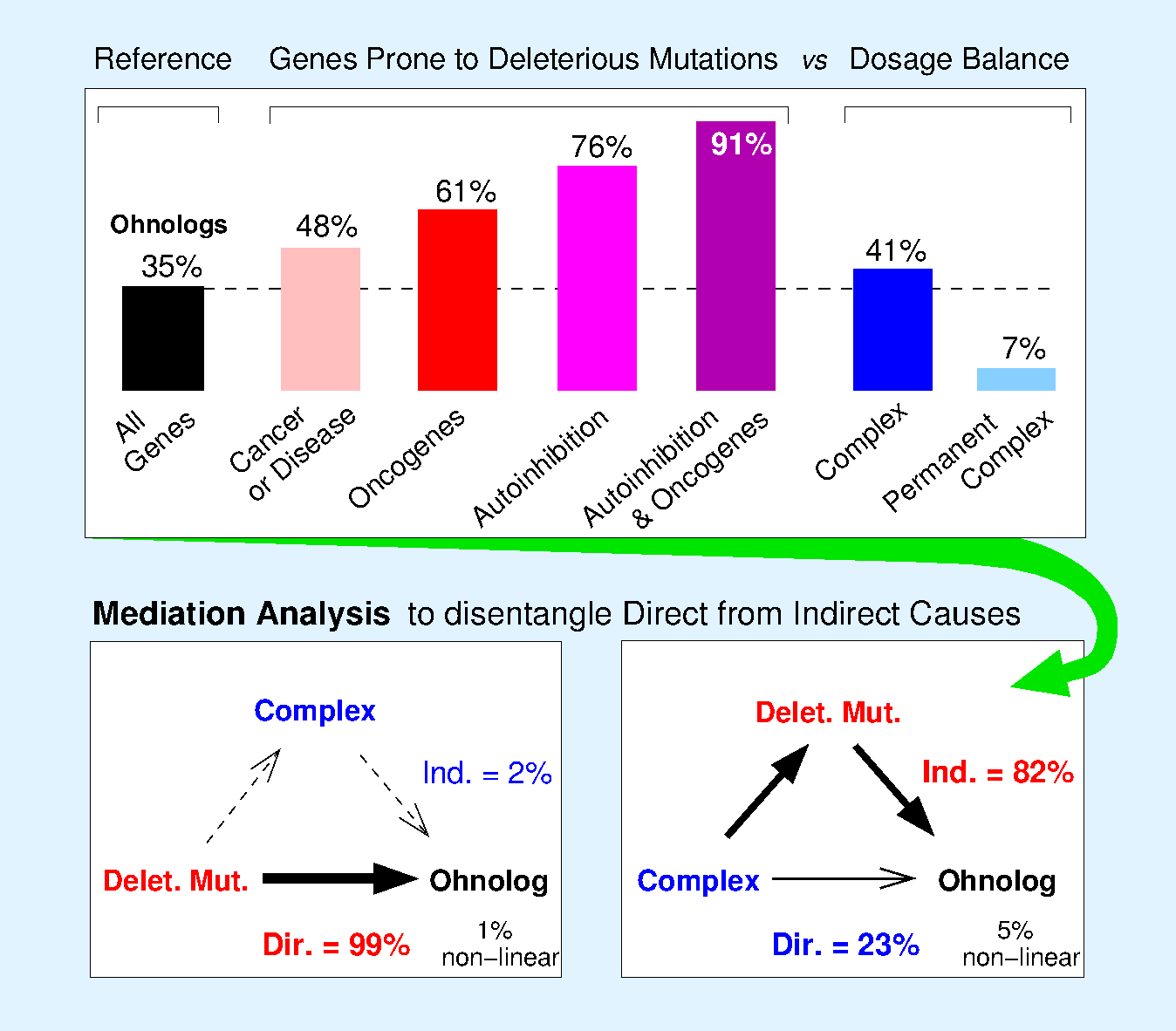
Figure 3.
Biased retention of ohnologs from whole genome duplication. Genes prone to dominant deleterious mutations, such as oncogenes and genes with protein autoinhibitory folds, have retained ohnologs from the two rounds of WGD in early vertebrates. By contrast, genes prone to dosage balance constraints have a mixed retention of ohnologs which can be accounted through indirect effect by quantitative Mediation Analysis disentangling direct from indirect causes (Singh et al. Cell Rep 2012).
|
| |
|
|
|
|
|
|
|
| |
|
|
This striking observation can be rationalized from a population genetics perspective by distinguishing WGD from SSD evolutionary scenarios sketched in Figure 4.
Hence, the enhanced retention of "dangerous" ohnologs, prone to dominant deleterious mutations, is shown to be a consequence of WGD-induced speciation and the ensuing purifying selection in post-WGD species, Figure 4.
This is further supported by stochastic simulations emphasizing the effect of small population sizes on the retention of "dangerous" SSD duplicates as well, Figure 5.
|
|
|
|
|
| |
|
|
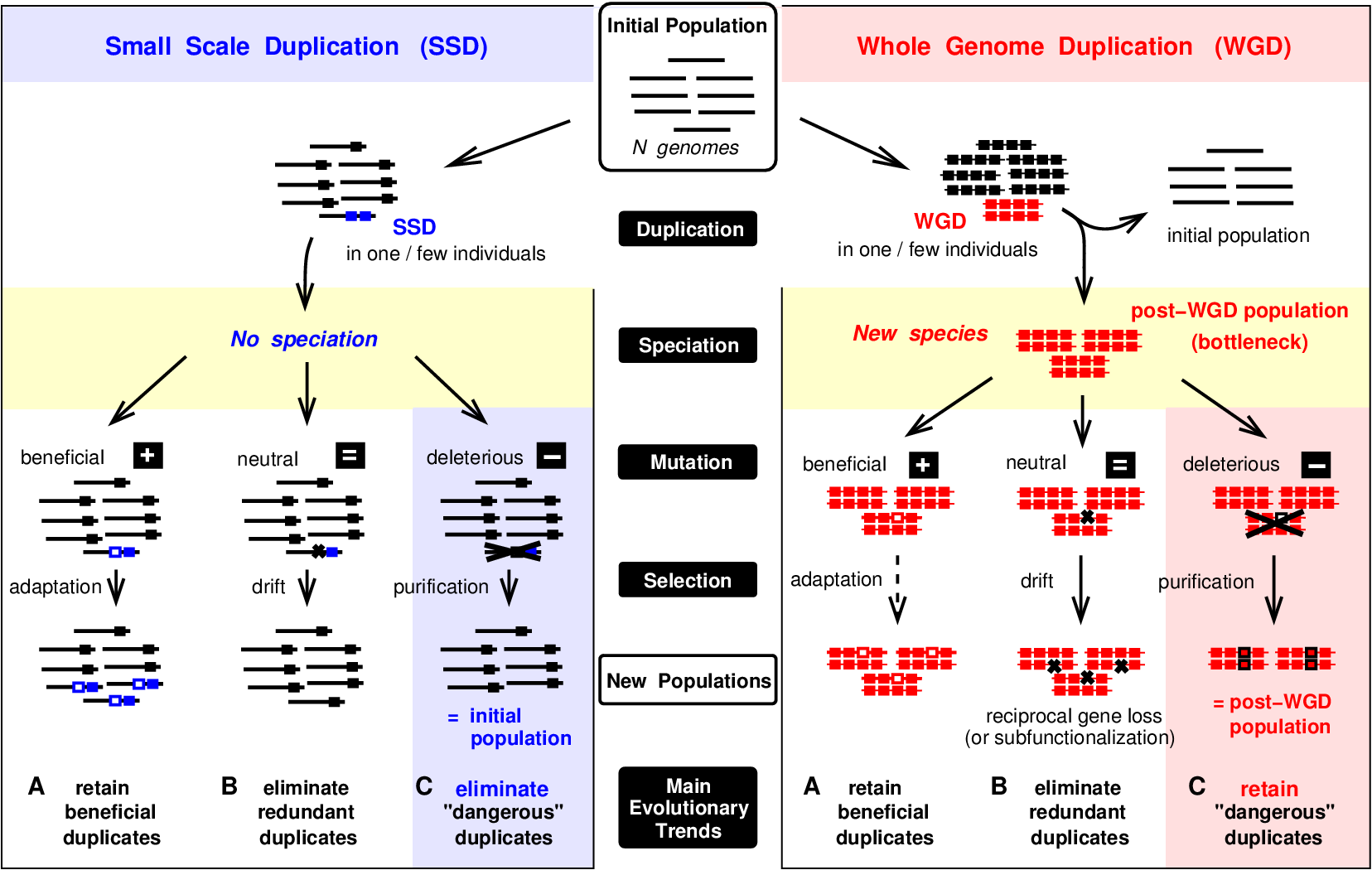
Figure 4.
Small Scale Duplication (SSD) versus Whole Genome Duplication (WGD) scenarios. Sketch of population genetics model to account for the different retention biased following SSD versus WGD scenarios (Singh et al. Cell Rep 2012).
|
|
|
|
|
| |
|
|
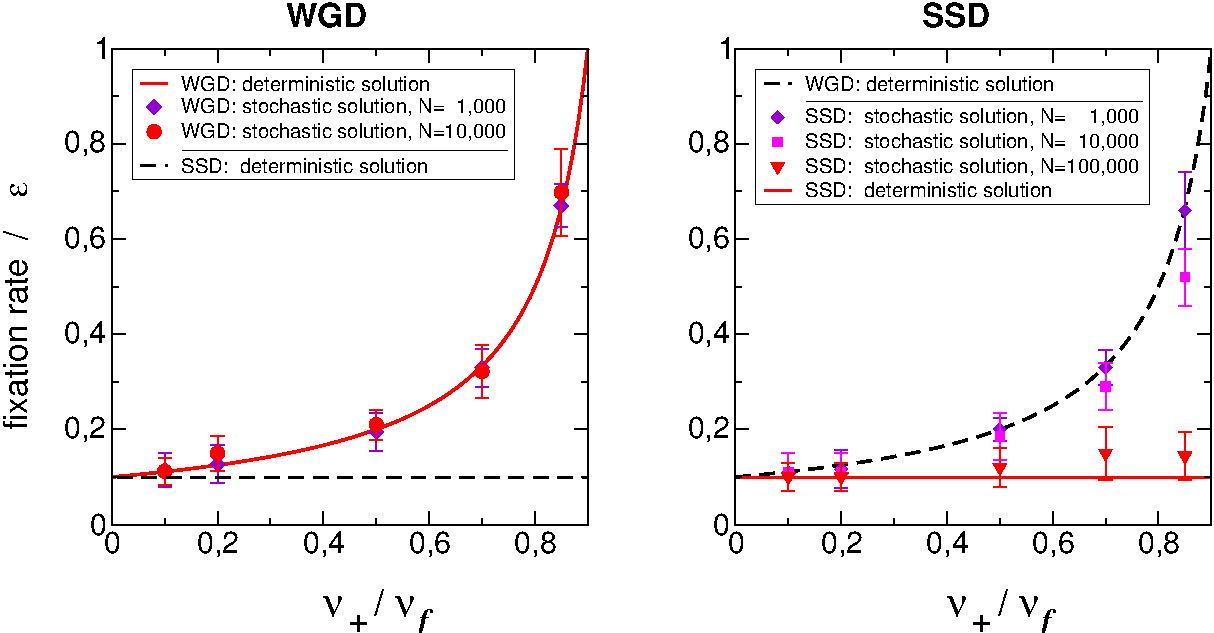
Figure 5.
Stochastic simulations of population genetics models for WGD versus SSD scenarios. Genes prone to dominant deleterious mutation (x-axis) are more likely to retained ohnologs from WGD than duplicates from SSD, unless the population size is small enough to bypass the adaptive fixation of SSD
(Malaguti et al. Theor Popul Biol 2014).
|
| |
|
|
|
|
|
|
|
| |
|
|
Yet, it is important to emphasize that the retention of "dangerous" ohnologs from ancient WGD is only a stochastic though significant bias, as illustrated in Figure 6. While only 10% of genes have retained an ohnolog at each round of WGD, this fraction doubles to about 20% for "dangerous" genes, such as oncogenes, implying that the most likely evolutionary outcome in most but extreme cases is the loss of ohnolog copies (e.g. 90% versus 80% of lost ohnolog copies for all genes versus oncogenes).
|
|
|
|
|
| |
|
|
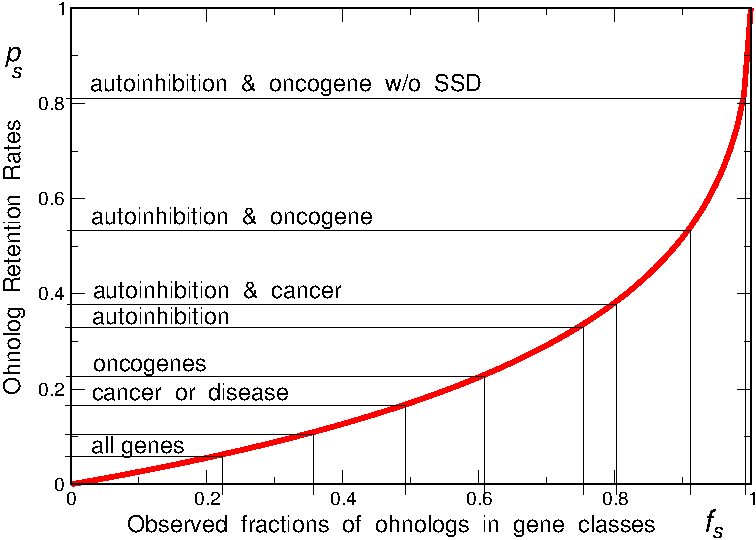
Figure 6.
Fraction of retained ohnologs at each round of whole genome duplication.
Note hat the most likely evolutionary outcome in most but extreme cases (such as oncogenes with autoinhibitory protein folds and no SSD copy) is the loss of ohnolog copies (e.g. 90% versus 80% of lost ohnolog copies for all genes versus oncogenes).
|
| |
|
|
|
|
|
|
|
| |
|
|
More recently, we have extended these correlation and mediation analyses to more than two or three genomic properties, using our information-theoretic method for network reconstruction (miic), Figure 7.
The resulting causal network, predicted by miic, relates the origin of duplicated genes in the human genome (i.e. ohnolog, SSD or CNV gene duplicates) to their genomic properties and association to diseases, Figure 7C. The reconstructed network implies that the retention of ohnolog duplicates is more directly linked to their susceptibility to dominant mutations and protein autoinhibitory folds than other genomic properties such as dosage balance constraints in protein complexes, gene essentiality or expression levels, which do not exhibit direct links to ohnolog retention, Figure 7C. Hence, miic analysis based on observational data provides an independent confirmation as well as significant extension of our earlier findings and simple population genetic models.
|
|
|
|
|
| |
|
|
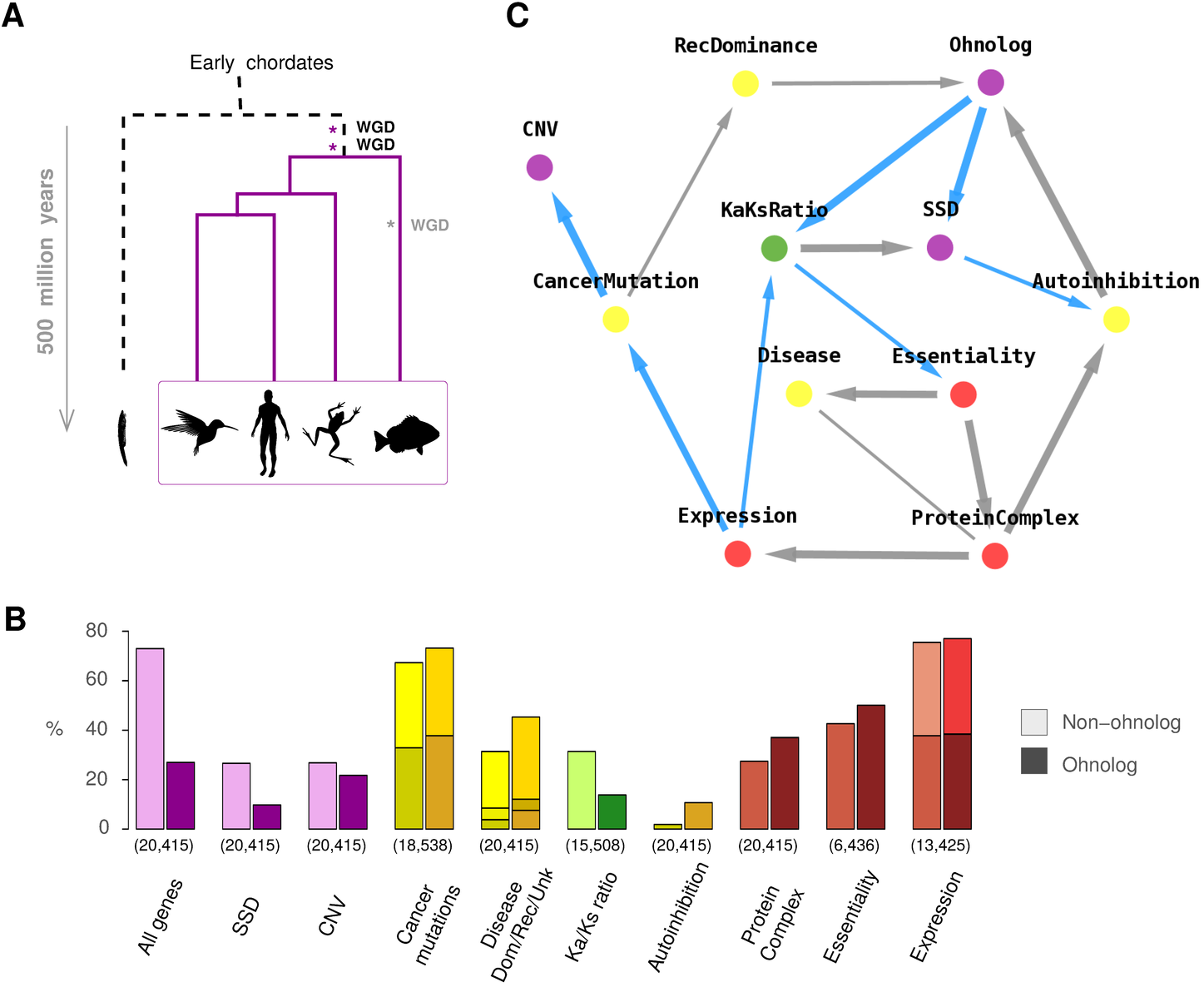
Figure 7.
MIIC network reconstruction between human genomic properties.
(A) Two rounds of whole genome duplication (WGD) have led to the evolutionary radiation of vertebrates (and similarly with a third 300-MY-old WGD in teleost fish). (B) Biased distributions of genomic properties within non-ohnolog and ohnolog genes retained from WGDs in early vertebrates. Numbers in brackets indicate the numbers of genes for which each property is identified. (C) Genomic property network of human genes predicted by miic (blue edges correspond to repressions), Verny et al. 2017.
|
| |
|
|
|
|
|
|
|
| |
|
|
All together, these results support an evolutionary retention of ohnologs by purifying selection through dominant diseases in tetraploid species (consistent with the retention of ohnologs with low Ka/Ks ratio, Figure 7C, indicating sequence conservation) while small scale duplicated genes have been retained through positive selection (consistent with their higher Ka/Ks ratio, Figure 7C, indicative of underlying adaptation). These findings highlight the importance of WGD-induced non-adaptive selection for the emergence of vertebrate complexity, while rationalizing, from an evolutionary perspective, the expansion of gene families frequently implicated in genetic disorders and cancers.
|
|
|
|
|
|
|
|
|
| |
Related Publications
|
|
Verny L, Sella N, Affeldt S, Singh PP, Isambert H: Learning causal networks with latent variables from multivariate information in genomic data.
PLoS Comput Biol 13(10):e1005662 (2017).
Pubmed
| PLOS
| pdf
| supp
[recommended by F1000]
Singh P-P, Arora J, Isambert H:
Identification of ohnolog genes originating from whole genome duplication in early vertebrates, based on synteny comparison across multiple genomes.
PLoS Comput Biol, 11(7):e1004394 (2015).
Pubmed
| PLOS
| pdf
| supp
| Ohnologs server
Singh PP, Affeldt S, Malaguti G, Isambert H:
Human dominant disease genes are enriched in paralogs originating from whole genome duplication.
PLoS Comput Biol, 10(7):e1003754 (2014).
Pubmed
| PLOS
| pdf
Malaguti G, Singh PP, Isambert H. On the retention of gene duplicates prone to dominant deleterious mutations.
Theor Popul Biol, 93:38-51 (2014).
Pubmed
| ScienceDirect
| pdf
Affeldt S, Singh PP, Cascone I, Selimoglu R, Camonis J, Isambert H: Evolution and cancer: expansion of dangerous gene repertoire by whole genome duplications.
Med Sci, 29(4), 358-61 (2013) [french].
Pubmed
| EDPSciences
| pdf
Singh PP, Affeldt S, Cascone I, Selimoglu R, Camonis J, Isambert H:
On the expansion of "dangerous" gene repertoires by whole genome duplications in early vertebrates.
Cell Rep, 2(5), 1387-1398 (2012).
Pubmed
| CellPress
| pdf
| supp
[featured by Le Point, Biofutur, CNRS]
|









