|
|
|
|
Genome duplication in tumor progression and resistance to treatment Despite continued progress in tumor treatments, cancer has become the first cause of death in Western Europe, where it outnumbers cardiovascular disease. Genome heterogeneity of cancerous cells within tumors is at the root of cancer resistance to treatment and explains much of the remaining therapeutic limitations. In particular, recent findings from cancer genome sequencing projects suggest that nearly 40% of tumors underwent whole genome duplication at some point of tumorigenesis, yet its impact on cancer progression and drug resistance is only beginning to emerge. In this project, we analyzed gene and genome alterations in breast tumor datasets from the COSMIC database, using our information-theoretic method for network reconstruction (miic), Figure 1. |
||
|
|
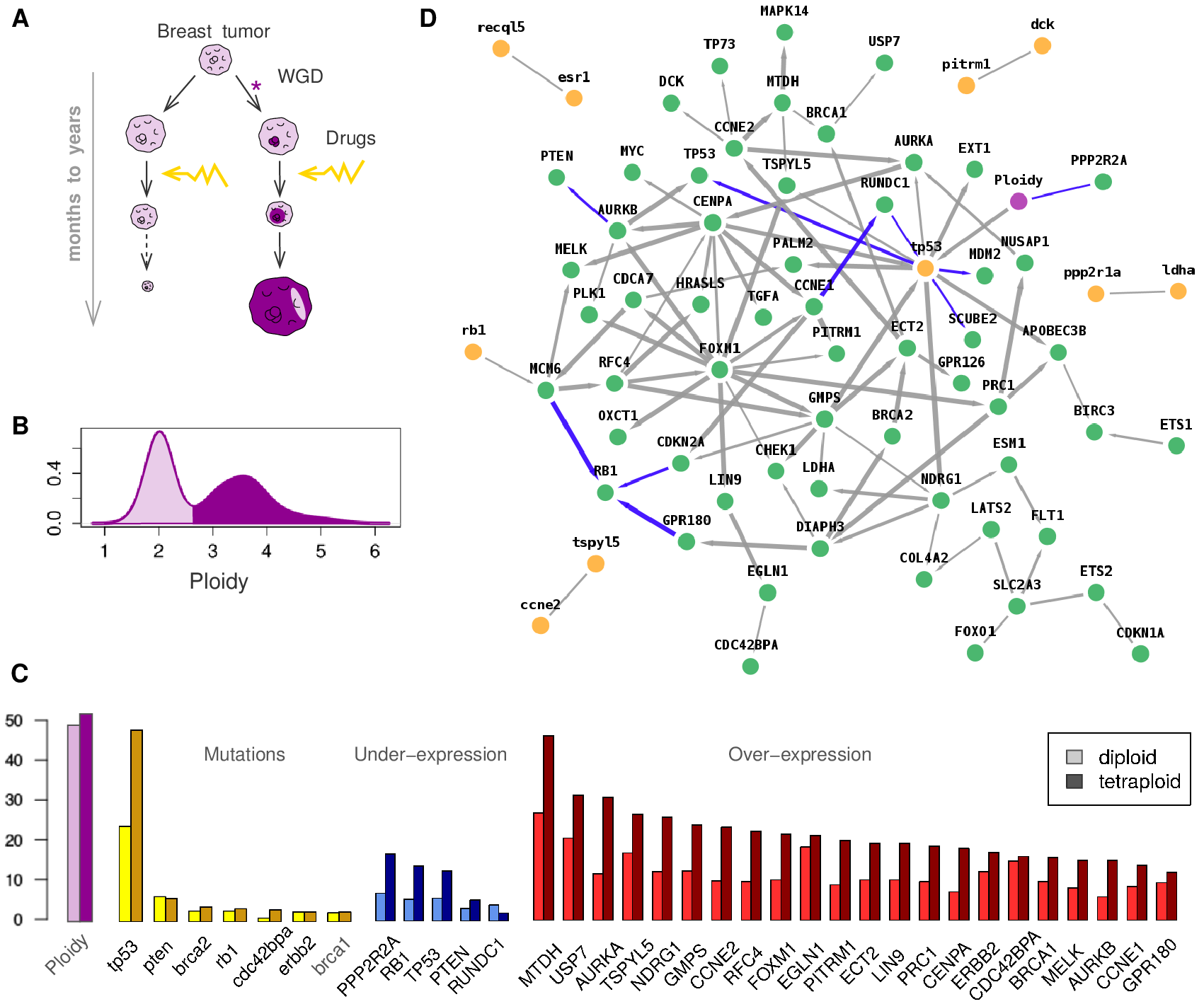
Figure 1. Genome duplication in tumor progression and resistance to treatment. (A) Tumor development and drug resistance in the presence of tetraploid tumor cells following whole genome duplication (WGD). (B) Ploidy distribution and (C) genomic alterations: ploidy, mutations, normalized under-expression and over-expression changes in 807 tumor samples from the COSMIC database. (D) Genomic alteration network obtained between average ploidy (violet), gene mutations (yellow, lower case) and under- or over-expressions (green, upper case). Blue edges correspond to repressions. Verny et al. PLoS Comput Biol 2017. |
||
|
The network predicted by miic is shown Figure 1D. Due to the limited numbers of samples (N=807) and recurrent gene mutants (Figure 1C), most gene mutations are not confidently linked to any altered expression levels, with the notable exceptions of TP53 and RB1 mutations, which have a significant impact on gene expressions, Figure 1D. Interestingly, the overall effect of tetraploidization on normalized gene expression, Figure 1C, is predicted to be largely indirect and mediated by TP53 mutations which lead to dysregulation of mitosis controling genes, such as the under-expression of PPP2R2A and over-expression of AURKA and CENPA genes. In addition, tetraploidy and TP53 mutations tend also to be concomitant with over-expression of metabolic (GMPS) and cell-growth modulating genes (TSPYL5, NDRG1 and FOXM1), favoring tumor progression and metastasis, as well as higher expression of APOBEC3B, which promotes mutational heterogeneity within tumors and, thereby, their drug resistance through subclonal selection. Hence, miic results provide a direct link between the long-known incidence of TP53 mutations in (breast) cancer and the tetraploidization of tumor cells. These results, supported by a number of recent reports, shed light on the poor prognosis associated with tetraploid tumors and their resistance to chemotherapy. This presumably occurs as tetraploid cells can exploit their genome redundancy and heterogeneity to evolve resistance strategies under drug treatments, Figure 1A.
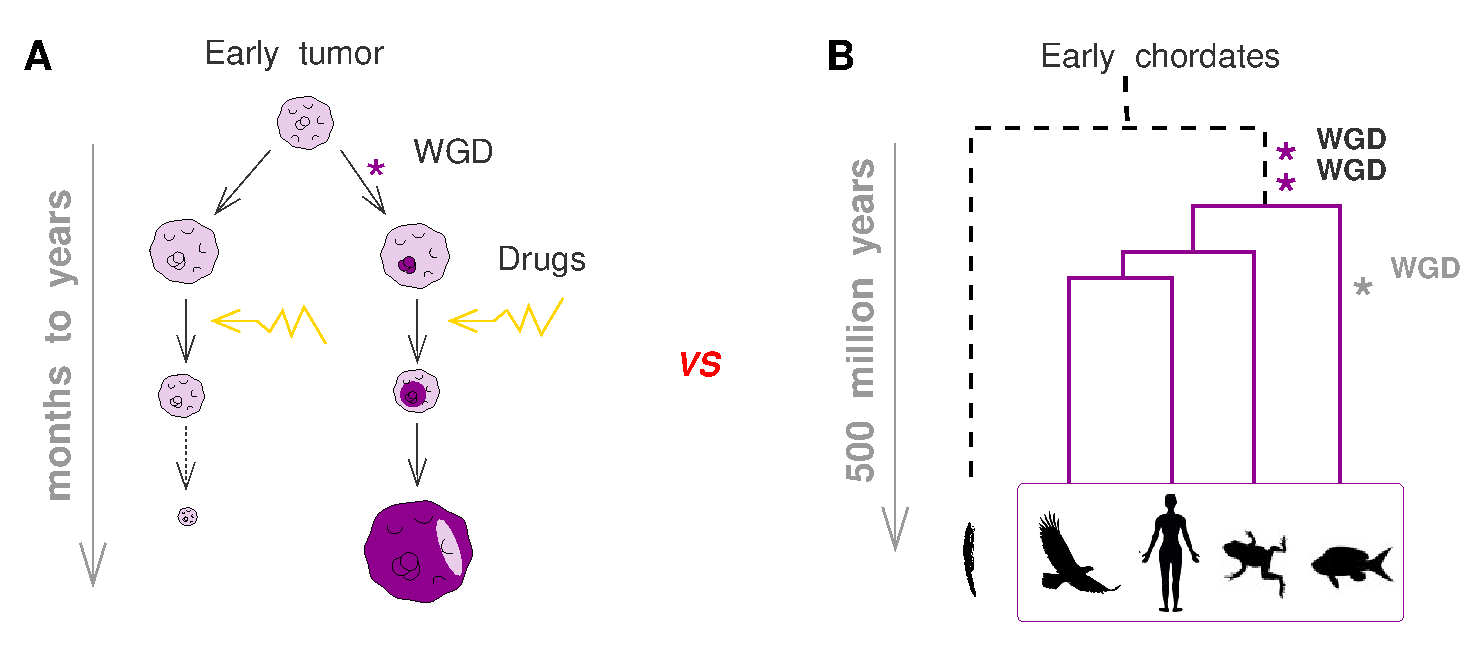
Interestingly, this dynamics of tetraploid tumors in the course of cancer progression and treatment, Figure 2A, echoes the success of tetraploid species in the course of eukaryote evolution, Figure 2B. Indeed, genome doubling events, possibly associated to environmental changes, have repeatedly led to successful evolutionary radiations of biodiverse species, such as the vertebrates and the flowering plants, see Figure 1 on the consequence of WGD in vertebrates.
|
|||
|
|
Figure 2. Role of genome duplication in tumor resistance and vertebrate diversity. (A) Tetraploid cells in tumors exploit their genome redundancy and heterogeneity to evolve resistance strategies under drug treatments. (B) Similarly, whole genome duplications, possibly associated to environmental changes, have repeatedly led to successful evolutionary radiations of biodiverse species, such as the vertebrates and the flowering plants. Adapted from Verny et al. PLoS Comput Biol 2017. |
||
|
|
|||
|
Related Publications |
|