|
|
|
|
Information maximization in mixed-type data from medical records The precise diagnostics of complex diseases, such as cancer and neurodegenerative diseases, requires to integrate a large amount of information from heterogeneous biomedical and clinical data, whose direct and indirect interdependence is notoriously difficult to assess. These diagnostics and related treatments should also take into account age-related comorbid medical conditions, such as diabetes, hypertension and cardiovascular diseases, which concern a large fraction of diagnosed patients, as the incidence of cancer and neurodegenerative diseases increases with age. These comorbid medical conditions influence treatment decisions as well as short- and long-term survival of patients but are often overlooked in clinical trials. This highlights the need to directly analyze real life medical records to uncover unsuspected associations and possible cause-effect relationships between various pathologies, specific treatments and any other relevant information recorded in the medical history of patients. To this end, we first developed and implemented an efficient machine learning approach to simultaneously compute and assess the significance of multivariate information in medical records, by maximizing information-theoretic measures including finite sample effects, Figure 1, between any combination of continuous or mixed-type variables in medical records. |
||
|
|
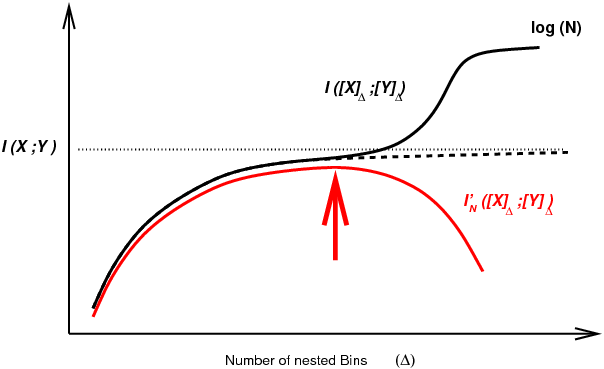
Figure 1. Mutual information computation between continuous or mixed-type variables for finite dataset. Arbitrary discretization of continuous variables tends to underestimate mutual information for small number of bins, while overestimating it for large number of bins due to limited number of samples. Alternatively, local metric approaches have been proposed to estimate mutual information (Kraskov 2004), based on k-nearest neighbor (kNN) statistics. However, the statistical significance of kNN information estimates remains difficult to assess in practice, thereby limiting their use to uncover (conditional) independences between continuous or mixed-type variables from real-life datasets. To circumvent this difficulty, we developed and implemented an efficient machine learning approach to simultaneously compute and assess the significance of multivariate information through an optimum partitioning of continuous variable(s) (red arrow) after introducing a complexity term to account for the finite size of the dataset, Cabeli et al (2020). |
||
|
|
|||
|
|
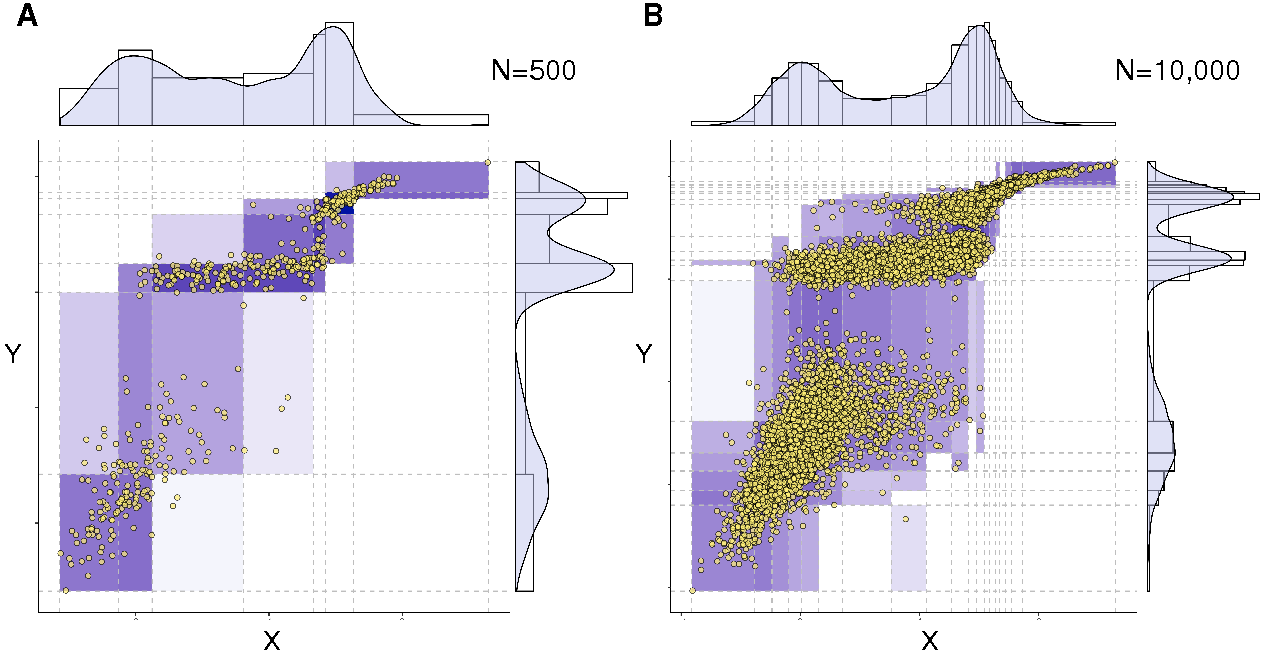
Figure 2. Optimum bivariate discretization for mutual information estimate. The information-maximizing partition yields (A) I(X;Y)=1.042 for N=500 samples and (B) I(X;Y)=1.165 for N=10,000 samples, as compared to the exact expected value I(X;Y)=1.2, Cabeli et al (2020). |
||
|
|
|||
|
|
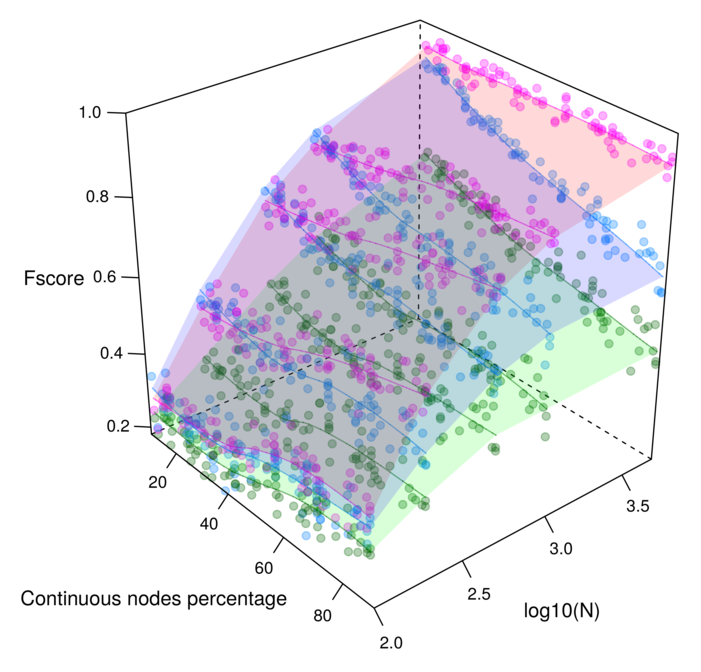
Figure 3. Reconstruction of benchmark networks for mixed-type non-gaussian datasets. F-scores obtained for 100 node random networks with average degree 3 reconstructed from N=10-10,000 samples. The parameter free Miic results (purple) are compared to alternative existing methods for mixed-type data: MXM (green) including one ajusting significance parameter (alpha) and CausalMGM (blue) including one ajusting regularization parameter (lambda). Cabeli et al (2020). |
||
|
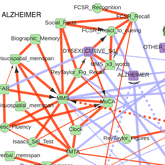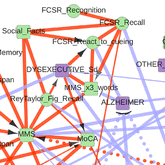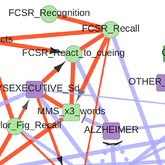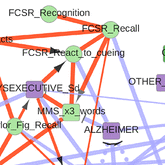
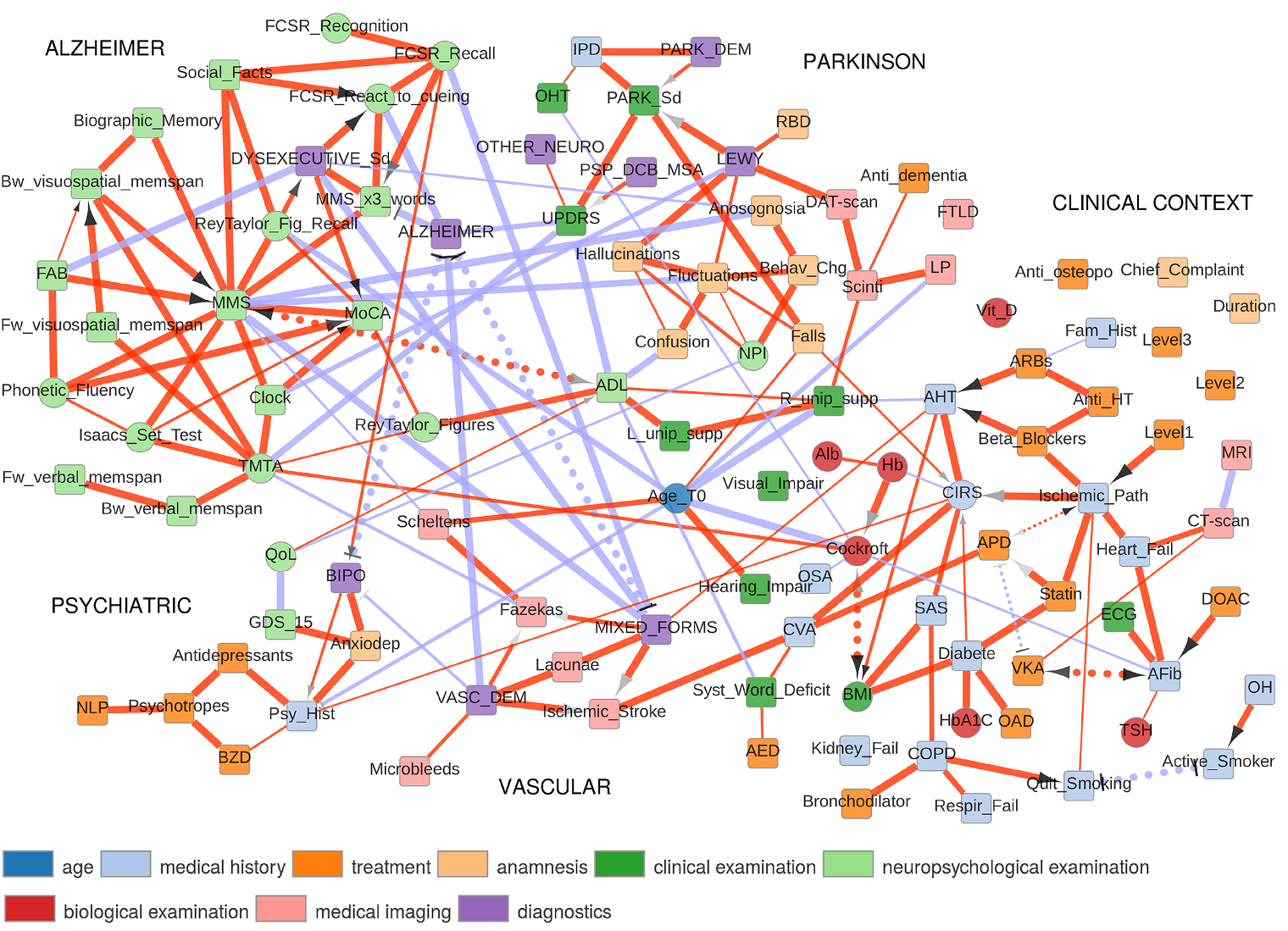
The first application concerns the reconstruction and visualization of a clinical network obtained from analyzing the medical records of 1,628 elderly patients consulting at La Pitie-Salpetriere hospital, Paris, for cognitive disorders (Collaboration with Prof M Verny). The dataset contains 107 variables of different types (ie 19 continuous and 88 categorical variables) and heterogeneous nature (ie variables related to previous medical history, comorbidities and comedications, scores from cognitive tests, clinical, biological or radiological examinations, diagnostics and treatments). The resulting graphical model, Figure 4, visually captures the global interdependencies in these medical records without any a priori hypothesis between clinically relevant and other variables. |
|||
|
|
Figure 4. Reconstruction and visualization of a clinical network from medical records of elderly patients with cognitive disorders. Collaboration with Prof M Verny, La Pitie-Salpetriere hospital, Paris (Cabeli et al 2020). |
||
|
The global analysis recovers well known relations as well as novel direct and indirect relations between specific variables in these medical records. In particular, we uncover a direct relation between the demyelinization and axonal loss caused by cerebrovascular accidents (fazekas scale) and the atrophy of the hippocampus (Scheltens scale). This provides some physiological insights linking the consequence of vascular accidents to the atrophy of important brain structures associated to cognitive impairment. |
|||
|
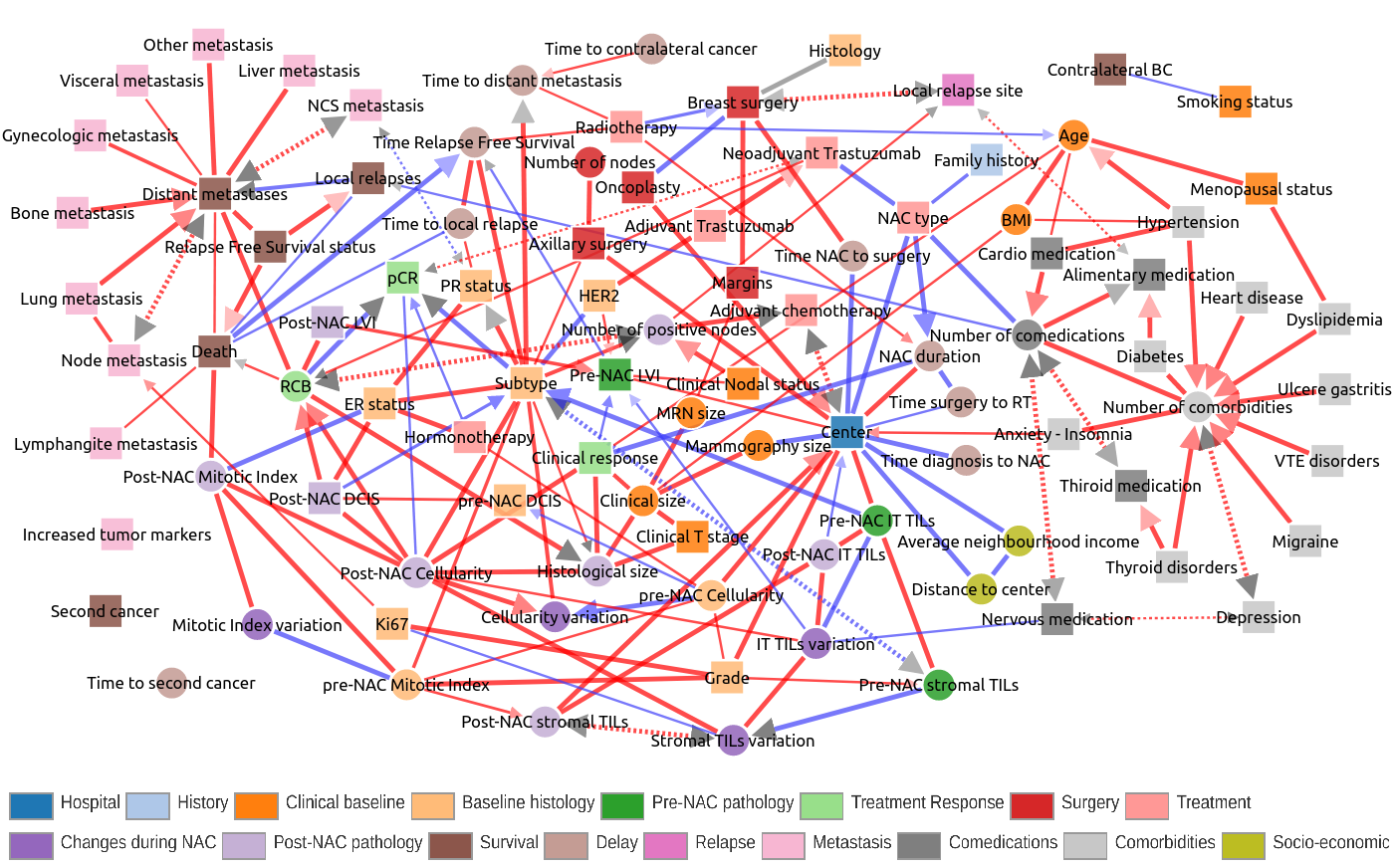
In particular, Figure 5 recapitulates the typical clinical follow-up of breast cancer patients from early diagnosis to successive treatments and recovery or relapses, metastasis. Yet, unexpected associations and biases were also uncovered through the clinical network analysis, such as i) the significant effects of the hospital center (Paris or St Cloud) on patient clinical profile at diagnosis and subsequent clinical practice or ii) the importance of the variation of TILS responses to treatment on the prognosis of the disease. |
|||
|
|
Figure 5. Interactive exploration of a global clinical network from a large breast cancer cohort treated by neoadjuvant chemotherapy. Collaboration with Prof F Reyal, Institut Curie, Paris (Sella et al 2022). |
||
|
However, MIIC predicts that pCR is not directly linked to survival. Instead, the continuous RCB score is found to be a more informative prognosis marker of patient eventual death through distant metastases. Indeed, our analysis based on information maximization principles leads to the definition of a new unsupervised classification of RBC scores into three categories: low-RCB with very good to good prognosis, depending on cancer subtypes (HER2+, 0% death; HER2- ER/PR+, 0%; TNBC 5%; All subtypes, 2.6%), medium-RCB with good to poor prognosis (HER2+, 1.9% death; HER2- ER/PR+, 7.6%;TNBC 29.7%; All subtypes, 14.9%) and high-RCB with poor to very poor prognosis (HER2+, 28.6% death; HER2- ER/PR+, 23.7%;TNBC 66.7%; All subtypes, 40.3%). In particular, our analysis with optimum discretization of RCB scores implies that the pCR class corresponding to RCB=0 should not be distinguished from the low-RCB class excluding RCB=0, which has very similar death rates as the pCR class (RCB=0). This is in agreement with independent findings recently published by Symmans et al J Clinic Oncol 2017. Hence, our analysis suggests to use a more informative, three-level classification of Residual Cancer Burden (RCB) scores than the (too conservative) pathological Complete Response (pCR) criteria to improve the prognosis accuracy of breast cancer patients. In addition, MIIC suggests relevant combinations of predictive of prognostic biomarkers. MIIC may provide clues to combinations of new prognostic biomarkers likely to improve the prediction of response to chemotherapy, or post-NAC prognosis. Pre-NAC lymphovascular invasion (LVI) was found to be associated with both lower rates of clinical response and shorter relapse-free survival. Both RCB and post-NAC mitotic index, a parameter rarely used in practice but nevertheless reported to be a predictor of BC recurrence, appear to be strongly associated with the risk of death. MIIC may, therefore, be an efficient tool for identifying features likely to improve prognosis, by combining gold standard indicators with other parameters, such as post-NAC mitotic index, and post-NAC LVI, for example. More recently, MIIC has been further extended into a more reliable, interpretable and scalable causal discovery method, iMIIC, which brings a number of important improvements enhancing its causal discovery performance on large scale real-life datasets, such as 400,000 medical records of breast cancer patients from the Surveillance, Epidemiology, and End Results (SEER) program (Ribeiro-Dantas et al. 2024).
|
|||
|
|
|||
|
Related Publications |
Cabeli V, Verny L, Sella N, Uguzzoni U, Verny M, Isambert H: Learning clinical networks from medical records based on information estimates in mixed-type data
|