|
|
|
|
Self-assembly of bacterial RNA
Biological structures and processes, as well as their regulation,
rely on specific interactions between
biomolecular components of the cell. While the main functions of
protein-protein, DNA-protein and RNA-protein interactions have long been
recognized, the full extent of RNA-RNA interactions has only recently emerged with the discovery of numerous non-coding RNAs in living cells.
|
||
|
|
DsrA nanostructure predictions and PAGE evidences
|
||
|
|
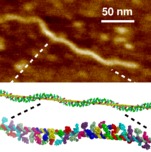
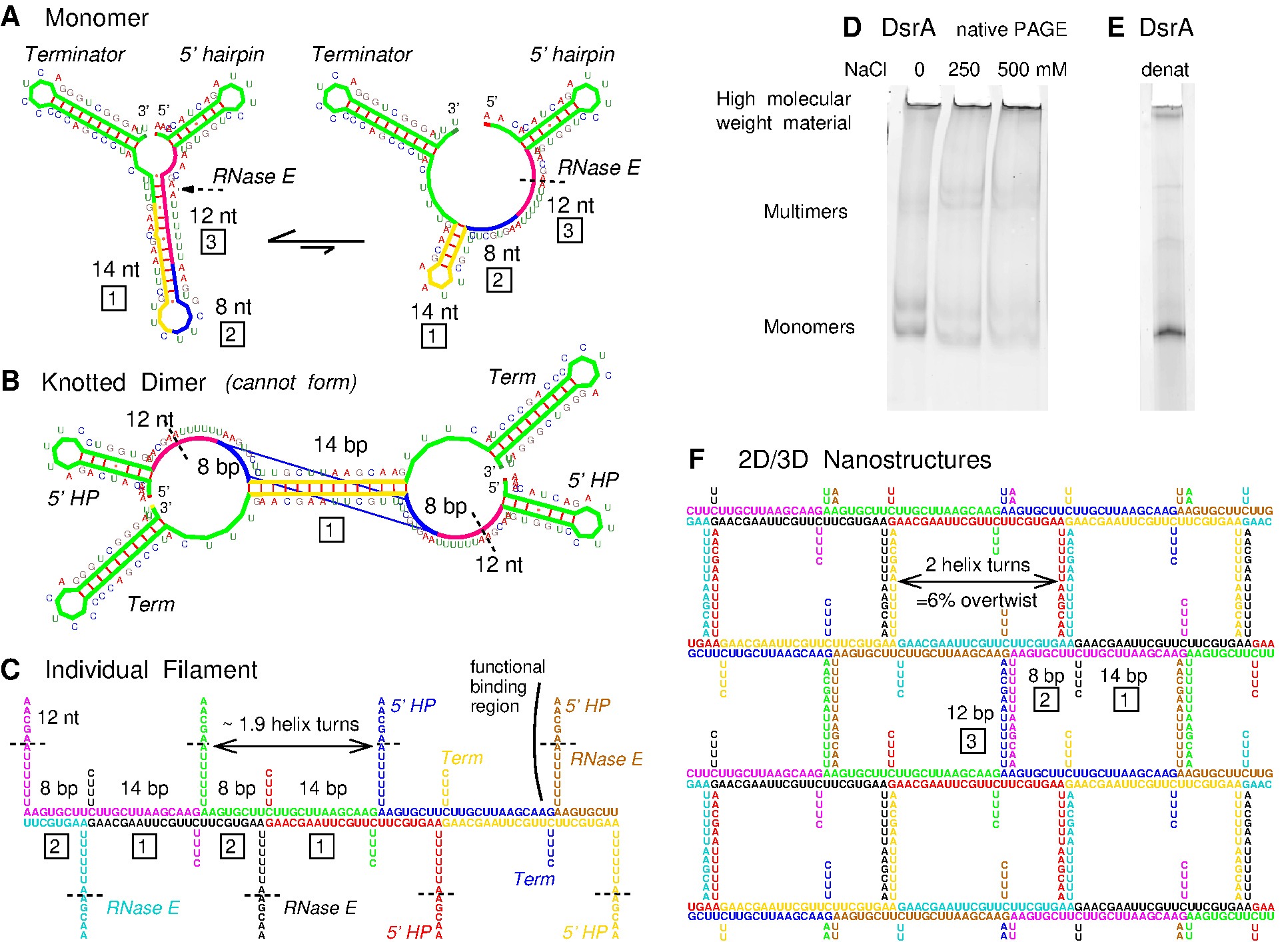
Figure 1. DsrA self-assembly prediction and PAGE evidence. (A) DsrA monomer structures with three contiguous self-complementary regions highlighted, 14-nt (yellow), 8-nt (blue) and 12-nt (magneta). (B) DsrA knotted dimer with both 14-bp and 8-pb duplexes cannot form for topological reasons. (C) Predicted DsrA filaments with coaxial stacking of 14-bp and 8-bp duplexes from successive self-assembled DsrA monomers (5' and terminator hairpins are not shown). DsrA functional binding region interacting with target mRNAs and Hfq protein and including an RNase E cleavage site (dashed line) is radially exposed from DsrA filaments. (D) Native PAGE of DsrA transcript under increasing salt concentration. (E) Denaturing PAGE of DsrA transcript. (F) Predicted nanostructure of laterally interacting DsrA filaments through many weak 12-bp bridging duplexes (5' and terminator hairpins are not shown). The main two regions with strongest self-complementary associations are an 8-nt region, 5'-AAGUGCUU-3', forming two central GU pairs, immediately followed by a second 14-nt region, 5'-CUUGCUUAAGCAAG-3', able to form a 14-bp Watson-Crick duplex. Yet, the contiguous locations of these two self-complementary regions along the DsrA sequence was expected to preclude the formation of fully paired DsrA dimers, with both 8-bp and 14-bp duplexes simultaneously formed, Figure 1B, because the corresponding knotted dimer cannot form for topological reasons. Instead, we anticipated that DsrA transcripts could in fact self-associate and form long double stranded RNA helices by coaxially stacking contiguous 8-bp and 14-bp duplexes from successive DsrA molecules, Figure 1C. Indeed, performing simple PAGE separations, we noticed that up to 30-50% of in vitro DsrA transcripts hardly migrate under typical electrophoretic conditions, Figure 1D. Although this high molecular compound is remarkably resistant to heat and urea denaturation, Figure 1E, we could rule out that it might come from read-through in vitro transcription using blunt PCR fragments for in vitro transcription by T7 or E. coli RNA polymerases. In addition, heat renaturation under increasing salt concentration was found to progressively convert low molecular weight bands (DsrA monomers, dimers,...) into the same hardly migrating band, Figure 1D. In fact, further analysis of DsrA middle region (in-between its 5'end hairpin and 3'end terminator hairpin, Figure 1A) shows that it also contains a 12-nt self-complementary sequence, which might possibly form a third duplex with two CU mismatches (Figure 1A), 5'-AACGAAUUUUUU-3'. While the corresponding binding energy, estimated as < 2 kcal/mol, is unlikely to yield isolated DsrA dimers in solution, the addition of many of these weak 12-bp duplexes is expected to induce efficient lateral interactions between individual DsrA filaments, self-assembled from the stronger 8-bp and 14-bp duplexes. The resulting DsrA 2D/3D nanostructures, Figure 1F, may thus provide strong local concentration enhancements enabling the formation of weakly paired 12-bp duplexes with possible implication for DsrA degradation and regulatory functions. The actual supramolecular structures of DsrA self-assemblies were then investigated using atomic force microscopy (AFM) and fluorescence microscopy techniques. |
||
|
|
AFM observation of robust filamentous self-assemblies
|
||
|
|
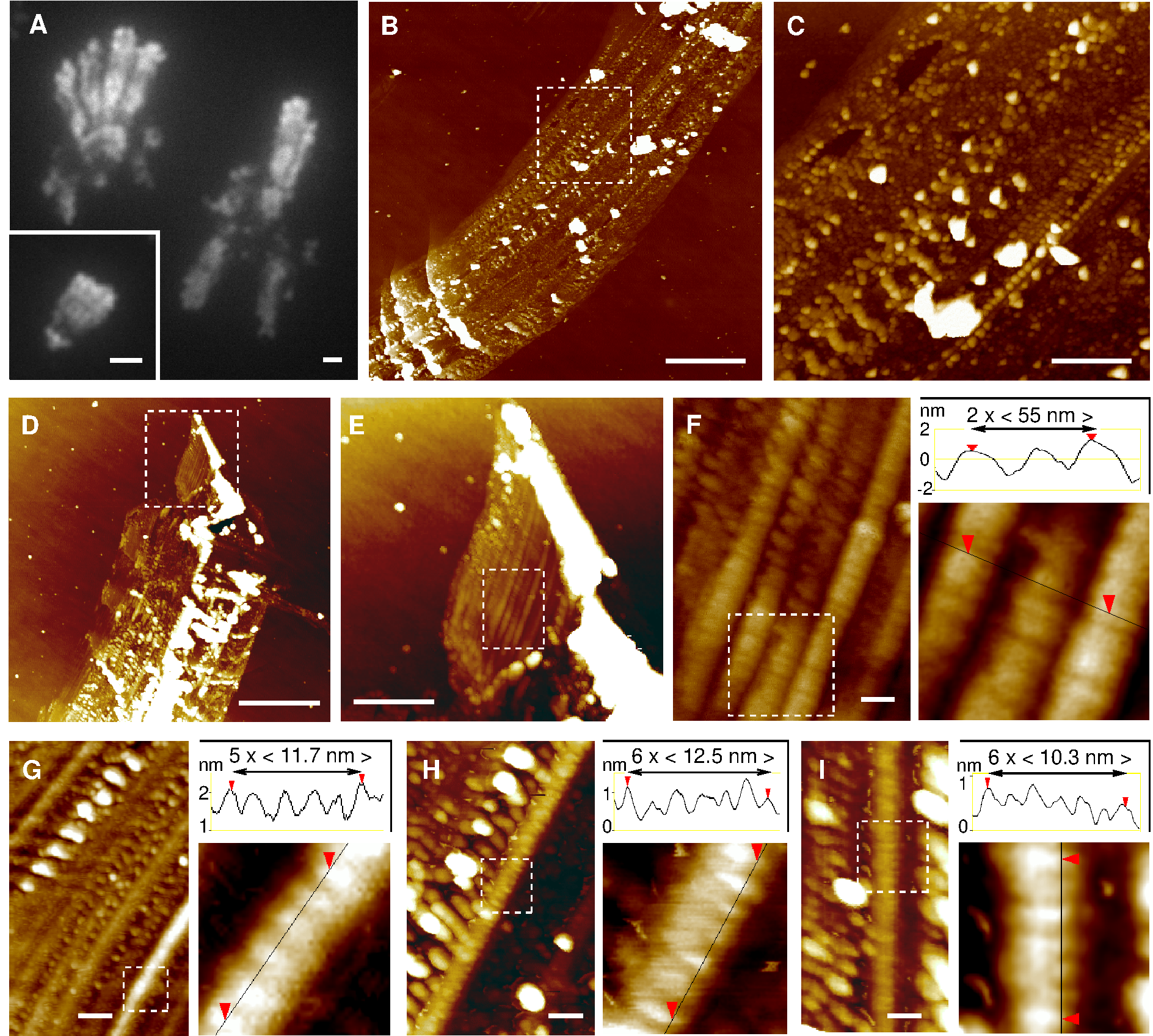
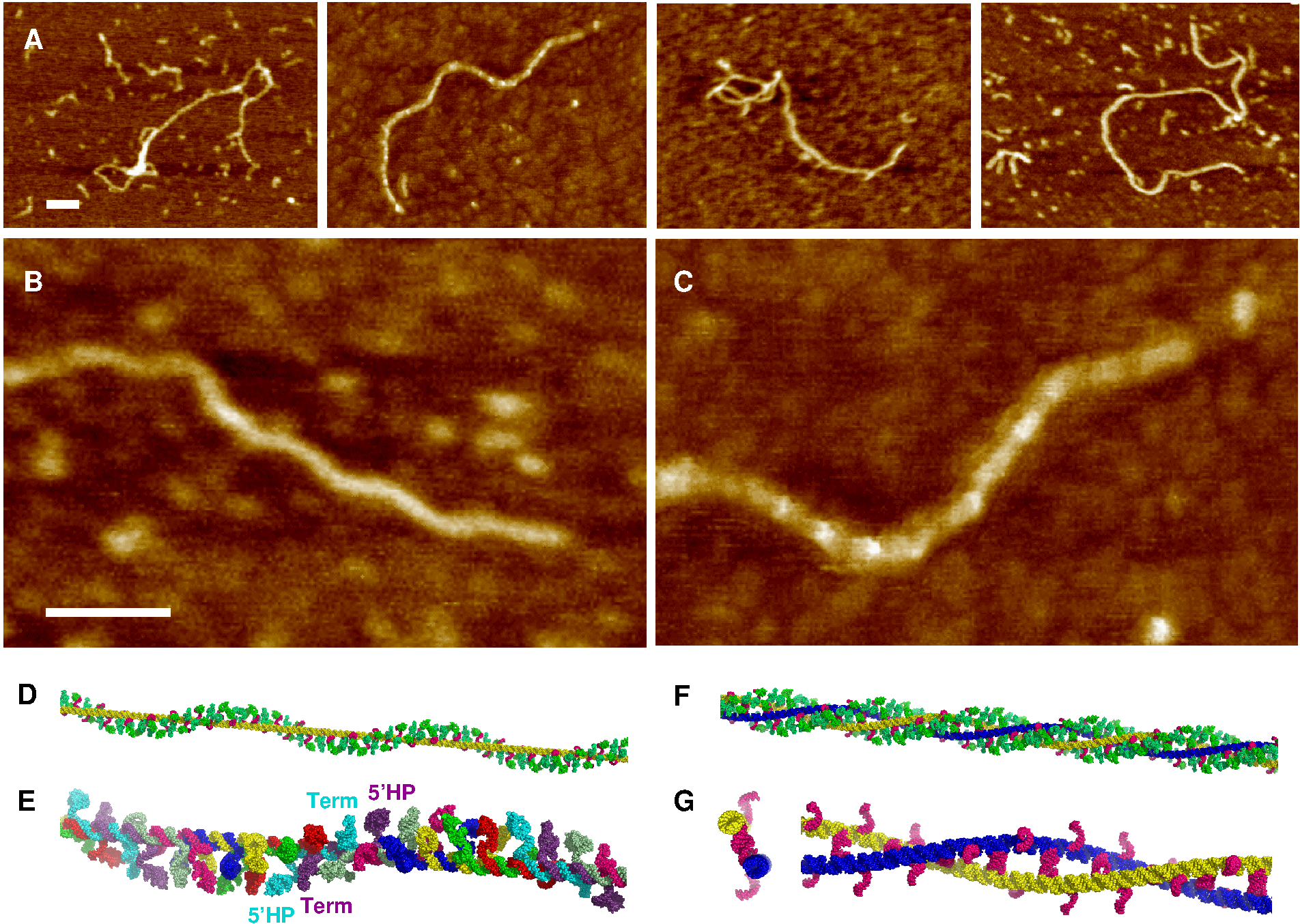
Figure 2. DsrA filament self-assembly. (A) AFM pictures of DsrA transcript: traditional dried sample preparation on mica. (B) , (C) Magnification of two characteristic longitudinal modulations with (B) 50 ± 5 nm period and (C) 23 ± 5 nm period. (D) Structural model using `pymol' of a single DsrA filament with a left-handed helical decoration from dangling 5' and 3' hairpins (green). (E) One left-handed helical turn (≈ 50 nm) with different colors for each successive DsrA molecule. (F) Structural model of two laterally interacting DsrA filaments (yellow and blue) with a left-handed helical decoration from dangling 5' and 3' hairpins (green). (G) Cross section (left) and one left-handed (super)helical turn (right) of two laterally interacting DsrA filaments with 12-bp bridging duplexes (magenta). Unpaired 12-nt self-complementary regions are radially exposed (magenta), hence, favoring further lateral accretion processes to occur and eventually larger DsrA self-assembly bundles to form, see main text. 5' and 3' hairpins are not shown for clarity. Scale bars: 50 nm. We also studied DsrA self-assembly by AFM in buffer, Figure 3. While single filaments and twisted filament bundles can be imaged in buffer, they do not adhere strongly to the surface or are destroyed after a few scanning images, as comparison between Figure 3 panels A and B shows. Yet, we observed some flat parallel filament bundles, which were found to adhere more tightly to the surface under buffer, than the more abundant helical filament bundles (Figure 3). These parallel arrangements of individual filaments could be imaged and analyzed in detail. Quantitative analysis of their height profiles as well as lateral distances between filaments, Figures 3C & D, largely confirm the proposed nanostructure model of parallel laterally interacting filaments, Figure 1F (see Figure 3 caption for details). |
||
|
|
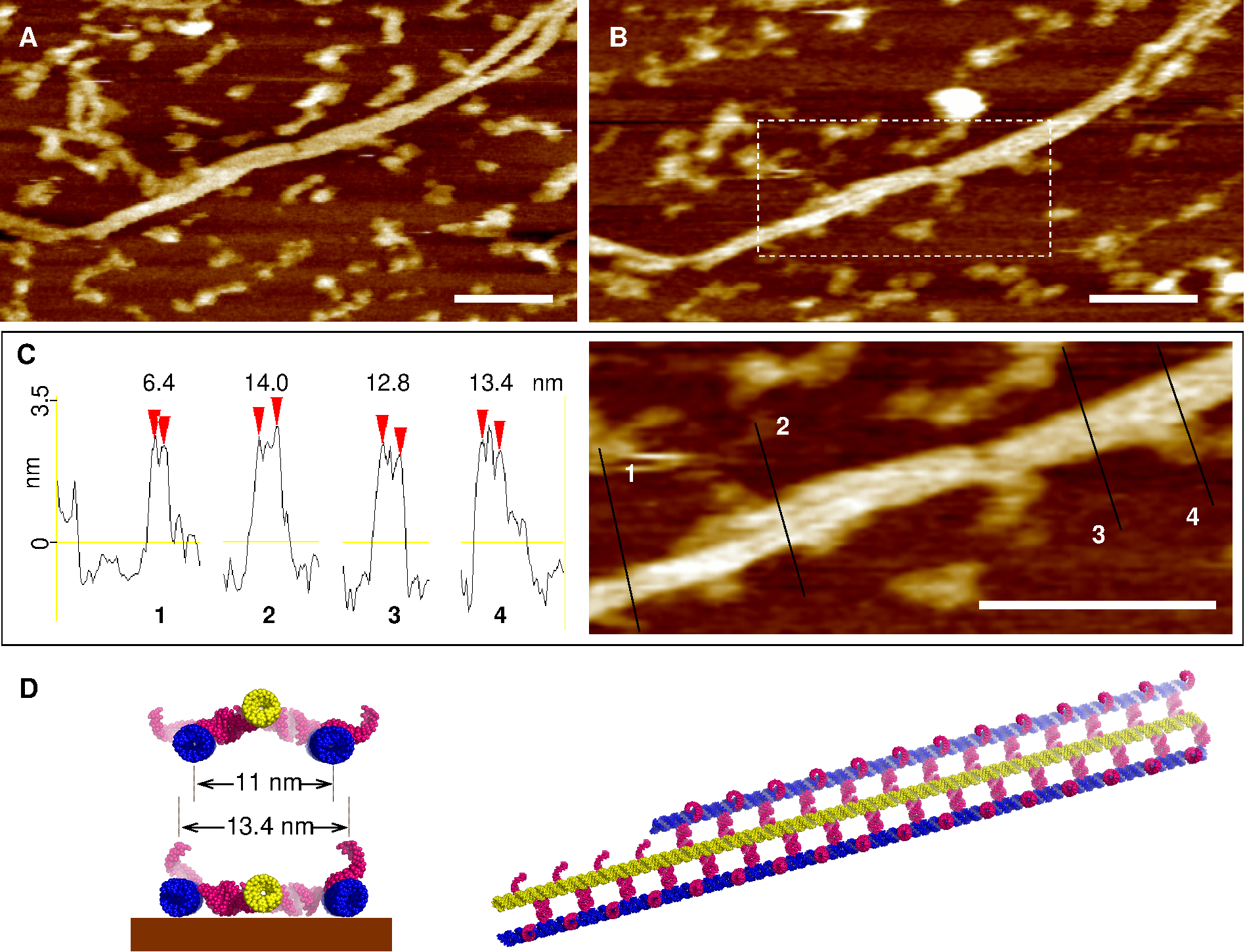
Figure 3. AFM imaging of parallel, laterally interacting DsrA filaments in buffer. (A, B) Flat bundles of parallel, laterally interacting filaments are found to adhere more tightly to the surface than the more abundant single filaments and helical filament bundles, which are easily damaged by the scanning tip in buffer (e.g., the single filament in the top left corner of panel A is heavily damaged in panel B after just a couple of AFM images). Note, that the flat filament bundle is also locally desorbed by the scanning tip (compare the top right corner in panels A and B), see main text. Scale bars = 100 nm. (C) Quantitative height and width profile measurements of these flat parallel filament bundles: height, 3.2 ± 0.2 nm; lateral distance between parallel filaments, 6.7 ± 0.4 nm (i.e., 13.4 ± 0.8 nm measured for three tightly adsorbed parallel filaments, as compared to the ≈ 11 nm for the undeformed predicted structure, panel D). Scale bar 100 nm. (D) Corresponding model of tightly adsorbed parallel filament bundle, see main text. The 5' and terminator hairpins are not shown but presumably account for the heights of the profiles (C), which are about 25% higher than a single dsRNA helix diameter (≈ 2.6 nm). Further evidence demonstrated, however, that these robust bundles of DsrA filaments actually resulted from the mechanical disruption of much larger but fragile anisotropic DsrA nanostructures during AFM sample preparation. |
||
|
|
DsrA forms extended but fragile nanostructures
In order to observe the structural organization of these large self-assemblies at a finer spatial resolution than fluorescence microscopy, we adopted a less disruptive AFM sample preparation by letting the DsrA transcript solution slowly dry (15-20 min) in between a silicon wafer and a glass coverslip. Indeed, direct fluorescent observations of the adsorbed nanostructures upon dewetting show that their characteristic micron scale pattern (Figure 4A) is left intact by the receding meniscus. This was confirmed by imaging the dry samples under AFM tapping mode, which show the same large, elongated structures, Figures 4B & D, as observed in solution under fluorescence microscopy (Figure 4A), while allowing for progressive zooming and local analysis of the fragile disrupted nanostructures under surface adhesion. |
||
|
|
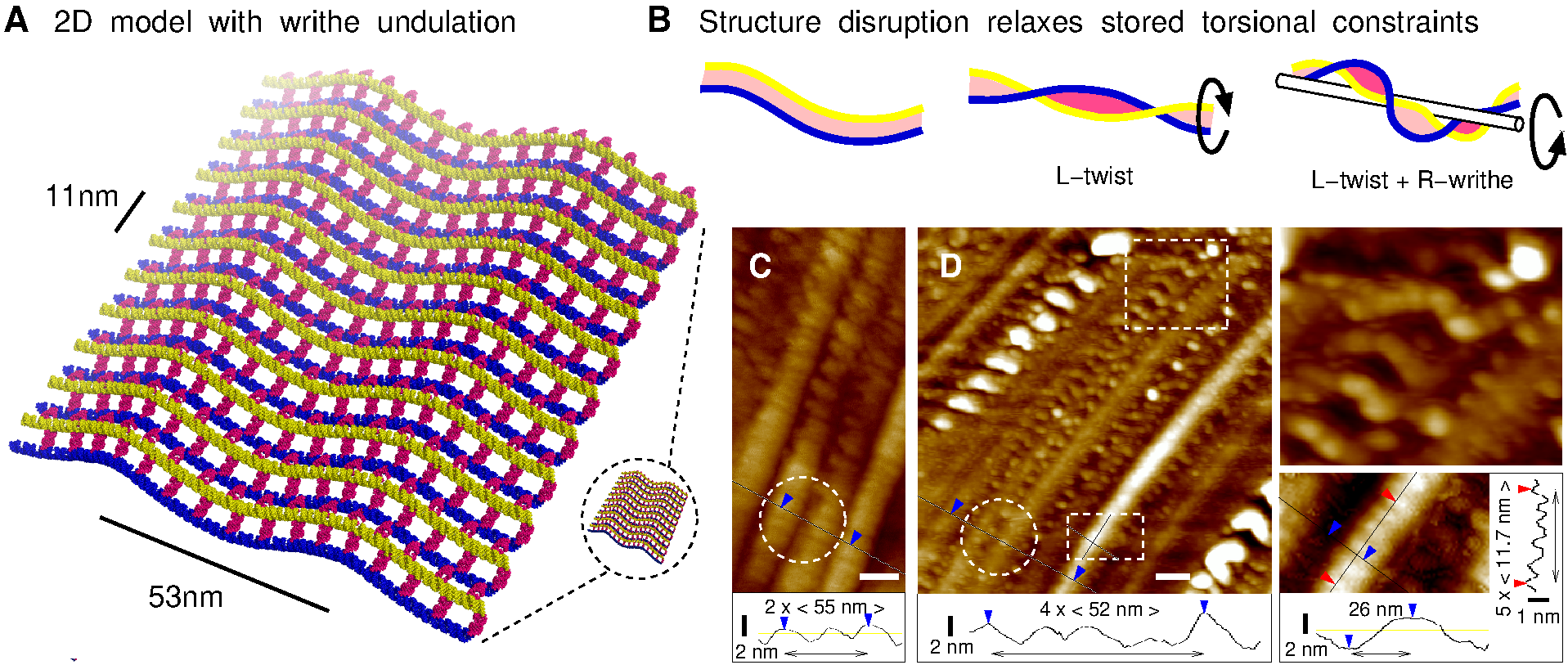
Figure 4. DsrA 2D/3D nanostructures. (A) Fluorescence microscopy pictures of DsrA self-assembly with fluorescent DsrA transcript from T7 RNAPol in vitro transcription (with 5% Alexa-488 labelled rUTP) or with DsrA non-fluorescent transcript from E. coli RNAPol in vitro transcription stained by SYBRgreen II (inset). Scale bars 2μm. (B-C) AFM pictures of similar disrupted DsrA nanostructures on silicon wafer substrate. The long direction of the supramolecular self-assembly is regularly interrupted by perpendicular coiled-up aggregates of various sizes, from micrometer range (Figure 4B bottom, also seen by fluorescence, Figure 4A) to 300-nm range (Figure 4C bottom) and even 30-nm range (Figure 4F, top and Figures 4G-I). Such self-similar-looking (or fractal-like) pattern at different length scales is typically characteristic of an underlying accretion mechanism. (D-F) By contrast, the least disrupted regions of these large DsrA nanostructures correspond to a regular 54 ± 5-nm striped pattern, parallel to the elongated direction of the supramolecular self-assembly. (G-I) Undisrupted stripes display also a 11.5 ± 2 nm striation. Scale bars: (A, B & D) 2 μm, (C & E) 500 nm, (F-I) 50 nm. The most regular regions (Figure 4E) from DsrA extended nanostructures (Figure 4D) correspond to smooth 54 ± 5 nm striped patterns (Figure 4F), parallel to the elongated direction of the supramolecular self-assembly. These smooth longitudinal stripes also clearly display perpendicular striations. The striation period for undisrupted stripes is 11.5 ± 2 nm (see quantitative profiles in Figures 4G-I). However, under structural disruption, the ≈ 11 nm striations are observed to coarsen into thicker aggregates, while the underlying longitudinal stripes concomitantly disappear and fuse (see top of Figure 4F and Figures 4G-I). Further accretion process eventually leads to the emergence of regularly spaced coiled-up agregates of increasing sizes, from 30-nm range (Figure 4F, top and Figures 4G-I) to 300-nm range (Figure 4C bottom) and even micrometer range (Figure 4B bottom, also seen by fluorescence, Figure 4A). Such self-similar-looking or fractal-like pattern at different length scales is typically characteristic of an underlying accretion mechanism. These striking structural patterns and their deformations are in fact consistent with an underlying self-assembly of aligned, laterally interacting DsrA filaments, as initially proposed in Figure 1F and directly observed under buffer, Figure 2. Indeed, the structural model for extended nanostructures of laterally interacting DsrA filaments, Figure 5A, requires that the natural rotation between successive 12-bp bridging duplexes (Figures 2B, D & E) is eliminated through a torsional constraint stored in a ≈ 6% right-handed overtwist of individual DsrA filaments. Such a torsional constraint is expected to be converted for energy minimization into a topologically equivalent helical curvature (or `writhe', Fuller, 1971) with a 53 ± 5 nm period, as depicted in Figure 5A. This is analogous to the linear-supercoiled transition of dsDNA above ≈ 2% overtwist (Marko & Siggia, 1995). Thus, while interacting DsrA filaments are essentially parallel, their collective writhe modulation is expected to lead to a regular striped pattern in the perpendicular direction, Figure 5A. Note that this predicted writhe-induced corrugation is entirely constrained by the twist / writhe geometric conversion rule without any fitting parameter other than the reported 11.7 bp/turn pitch of dsRNA helix (Conte, 1997; Clore, 1985; Arnott, 1986). Twist / writhe conversion is thus the most likely structural feature underlying the observed 54 ± 5 nm longitudinal stripes in large, elongated nanostructures, Figures 4D-F. This is further corroborated by the fact that the predicted ≈ 11 nm striation between every second filaments (Figure 5A) is also clearly observed along undisrupted stripes (see quantitative profile measurements in Figures 4G-I and Figures 5C & D). |
||
|
|
Disruption of DsrA nanostructures relaxes stored torsional constraints
|
||
|
|
Figure 5. 2D/3D model of extended DsrA nanostructure. (A) Structural model of 2D DsrA self-assembly. Individual DsrA filaments (blue and yellow) interact laterally by forming many weak 12-bp bridging duplexes (magenta), which requires that their natural left-handed rotation (Figures 2D & E) is eliminated by an opposite ≈ 6% right-handed overtwist. This is, however, converted for energy minimization into a topologically equivalent helical curvature (or `writhe') with a 53 ± 5 nm period producing a regular striped pattern (perpendicular to the aligned DsrA filaments); 5' and 3' hairpins are not shown for clarity. (B) Relaxation of stored torsional constraint by disruption of regular DsrA nanostructures: broken pairs of interacting filaments relax into left-handed superhelical conformations, `L-twist' (Figures 2F & G) and further cluster into larger filament bundles while the global helical winding constraint for pairs of unbroken filaments produces a characteristic local modulation pattern, due to an opposite twist / writhe cancellation, `L-twist+R-writhe'. (C-D) Corresponding AFM observation showing different characteristic patterns consistent with the extended nanostructure model (A) and its mechanical disruption (B), see main text. These structural predictions (Figures 5A & B) are in good agreement with the different characteristic patterns observed at the onset and early steps of the fragmentation of large DsrA nanostructures, as depicted in Figure 5D. Although still in one piece, the extended nanostructure is invaded by internal `cracks' along its laterally interacting filaments. At the same time, local disruptions of 12 bp duplexes are transmitted across tens to hundreds laterally interacting filaments, along the direction of the ≈ 50 nm stripes, as observed in Figures 4G-I and Figure 5D. Such mesoscopic correlation length scales provide a direct structural basis for the observed fragility of DsrA nanostructures, due to their stored torsional constraints. In addition, the relaxation of built-in mechanical constraints stored in DsrA extended nanostructures (Figure 5B) is also consistent with their observed global fragmentation, leaving only few intermediate sized pieces. This is analog to the well-known fragmentation property of `tempered' (safety) glass, which also stores internal mechanical constraints and shatters, as a result, into small pieces without leaving (dangerous) intermediate sized pieces. |
||
|
|
Functional role of DsrA nanostructures ?
|
||
|
|
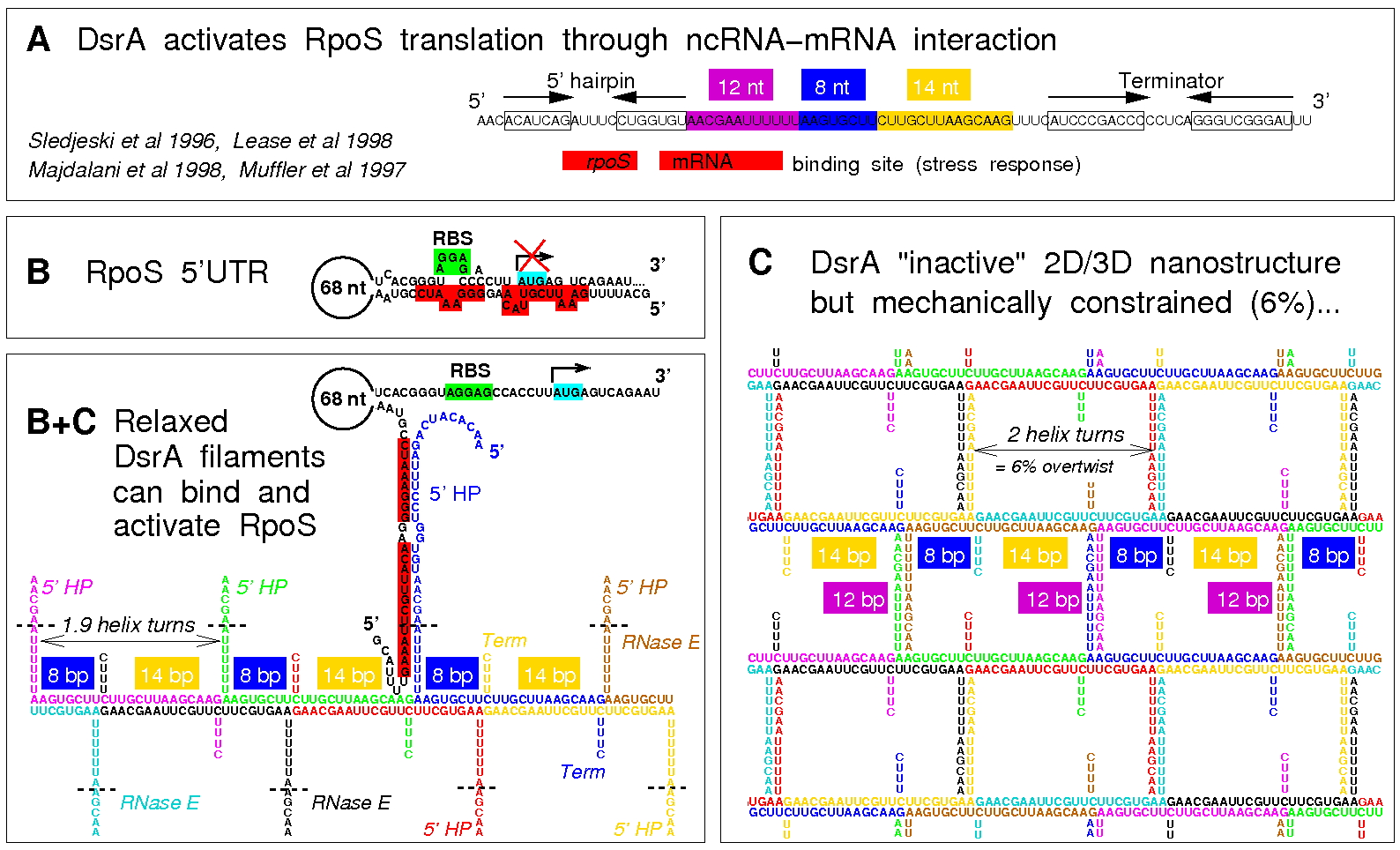
Figure 6. Working hypothesis for the functional role of DsrA nanostructures. (A) DsrA activates the translation of stress response sigma factor RpoS through ncRNA-mRNA interaction (binding site, red region). (B) 5'UTR of target mRNA rpoS highlighting the secondary structure fold sequesting the Ribosome Binding Site (RBS), thus preventing basal translation in absence of DsrA-rpoS RNA-RNA interaction. (C) Working hypothesis: the formation of the 2D/3D DsrA nanostructures enable to store large amount of DsrA ncRNA in a "functionally inactive" yet RNase resistant structural form (RNase E single strand binding site is rendered inaccessible through the formation of the weak 12-bp duplexes). (B+C) DsrA binding to its 5'UTR rpoS target requires the relaxation of stored torsional constraints through an induced structural switch of DsrA nanostructures triggering the sudden exposition of a large number of single-stranded functional binding sites, previously stored into the weakly paired 12-bp duplexes (B). We have investigated this functional scenario using in vitro transcription assays of fluorescent DsrA and/or target transcripts (Isambert, unpublished work). These results, Figure 7, demonstrate that the co-transcriptional assembly of DsrA leads in fact to metastable nanostructures at 37°C, Figure 7A, that can subsequently relax upon binding the target fluorescent-labeled 5'UTR of rpoS mRNA, Figure 7B, as expected from the working hypothesis proposed in Figure 6. In parallel to these in vitro analysis using FISH, we have also studied the formation and switching mechanism of these co-transcriptionally self-assembled DsrA nanostructures in vivo using FISH, Figure 7B. |
||
|
|
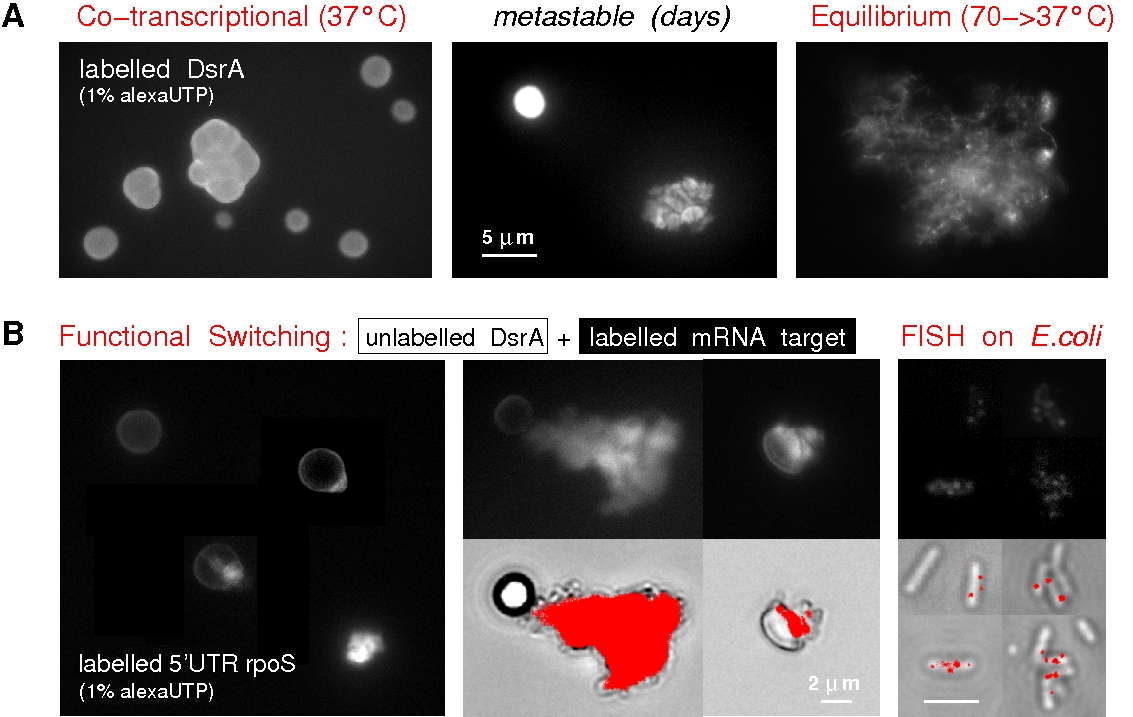
Figure 7. Experimental evidence of DsrA nanostructure regulatory function. (A) Co-transcriptional assembly of DsrA ncRNA in vitro leads to large metastable nanostructures (stable for days), which become easily disrupted under mechanical stress at 37°C or relax into mesh-like networks through thermal renaturation at 70°C. T7 in vitro transcription of fluorescent DsrA transcripts with 1% Alexa-488 labelled rUTP. (B) Functional switching mechanism can be demonstrated using unlabelled co-transcriptional assembly of DsrA nanostructures mixed to 1% Alexa-488 labelled 5'UTR rpoS binding region. 5'UTR target can only bind to mechanically relaxed regions of metastable DsrA nanostructures, hence, validating the proposed the "functionally inactive" structure and regulatory switch mechanism of DsrA co-transcriptional nanostructures. Co-transcriptionally self-assembled DsrA nanostructures can also be observed in vivo using FISH (Isambert, unpublished results). All in all, these results suggest that DsrA anisotropic nanostructures actually provide a useful platform for the control of DsrA regulatory functions under stress. In particular, the relaxation of stored torsional constraints through an induced structural switch of DsrA nanostructures was shown to trigger the sudden exposition of a large number of single-stranded functional binding sites, Figure 7B, previously stored as weakly paired 12-bp duplexes. Hence, the functions of DsrA nanostructures might in fact stem from the large-scale relaxation of their built-in structural constraints. |
||
|
|
|||
|
Related Publications |
|