| |
Main Peer-reviewed Publications
|
|
|
|
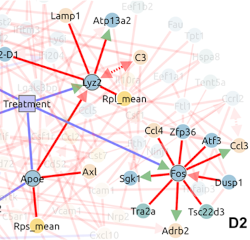
Manfroi B, Dang VD, Bui-Thi C, Jungmann A, Borzakian S, Tong Y, Beauvineau C, Dupuis L, Guffart E, Borst K, Nguyen NT, Alvarez-Simon D, Chyzak G, Schaefer S, Gerlach RG, Frischbutter S, Hamann A, Salem Wehbe L, El Behi M, Luka M, Menager M, Jouneau L, Boudinot P, Jung S, Winkler TH, Liblau R, Marignier R, Isambert H, Prinz M, von Kries P, Specker E, Walter J, Mahuteau-Betzer F, Fillatreau S: Induced regulatory B cells stably expressing IL-10 cure CNS autoimmunity by targeting local myeloid cells
Immunity, in press (2026).
Pubmed
| DOI
| pdf
|
| |
|
|
|
|
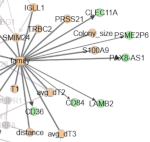
Donada A, Hermange G, Tocci T, Midoun A, Prevedello G, Hadj Abed L, Dupre D, Sun W, Milo I, Tenreira Bento S, Pospori C, Innes A, Willekens C, Vargaftig J, Michonneau D, Lo Celso C, Servant N, Duffy KR, Isambert H, Cournede PH, Laplane L, Perie L: Clonal memory of cell division in humans diverges between healthy haematopoiesis and acute myeloid leukaemia
Nature Commun, in press (2026).
Pubmed
| DOI
| bioRxiv
|
| |
|
|
|
|
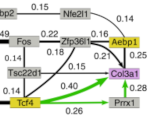
Fusilier Z, Clement A, Simon F, Calvente I, Jean-Marie R, Mathieu M, Calmettes V, Piastra-Facon F, Quintana-Perez Y, George Clement J, Crestey L, Lumineau E, Henninger R, Tonani M, Manriquez V, Lacerda L, Bensaid L, de Villemagne P, Piaggio E, Semetey V, Coscoy S, Martini E, Scita G, Gelly J-C, Ivaska J, Isambert H, Goudot C, Pierobon P, Lennon-Dumenil A-M, Moreau HD: Macrophages restrict tumor immune infiltration by controlling collagen topography
Science Immunology, 11 (117):eadw8291 (2026).
Pubmed
| DOI
| bioRxiv
| pdf
|
| |
|
|

|
|
Buvat I, Aarntzen E, Badawi RD, Ballesta A, Bock C, Chuah C-N, Fisher J, Graff J-P, Ilan Y, Isambert H, Ivanov P, Izu LT, Koenen HJPM, Langevin H, Lewis DY, Pires M, Sundar LKS, Tawakol A, ter Horst R, van Cranenbroek B, Weigelin B, Cherry SR, Beyer T: Cross-Disciplinary Methodologies for Whole-Person Research - Insights from EMPOWER2024
npj Imaging, 4, 19 (2026).
Pubmed
| DOI
| pdf
|
| |
|
|
|
|
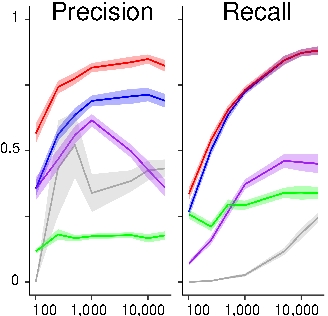
Lagrange N, Isambert H: An efficient search-and-score algorithm for ancestral graphs using multivariate information scores for complex non-linear and categorical data
Proceedings of the 42nd International Conference on Machine Learning (ICML2025), PMLR 267:32164-32187 (2025).
Pubmed
| DOI
| ICML
| pdf
|
| |
|
|

|
|
Dupuis L, Debeaupuis O, Simon F, Isambert H: CausalCCC: a web server to explore intracellular causal pathways enabling cell-cell communication
Nucleic Acids Res, 53, W125 (2025).
Pubmed
| DOI
| pdf
|
| |
|
|
|
|
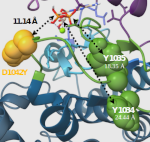
Jeanpierre M, Debeaupuis O, Brunaud C, Yancoski J, Riller Q, Hadjadj J, Stolzenberg M-C, Villarreal G, Katsicas MM, Villa M, Neves JF, Stephan J-L, Leonard C, Lazaro E, Ciron J, Boussard C, Mazerolles F, Magerus A, Pelle O, Masson C, Schmitt Y, Hoareau B, Vinit A, Neven B, Quartier P, Isambert H, Oleastro M, Danielian S, Parlato M, Rieux-Laucat F: In silico modeling guides identification of novel JAK1 variants associated with immune dysregulation
EMBO Mol Med, 17 (12), 3275 (2025).
Pubmed
| DOI
| pdf
|
| |
|
|
|
|
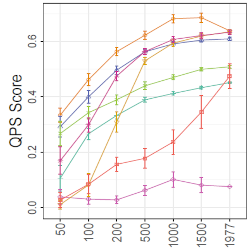
Sella N, Guinot F, Lagrange N, Albou LP, Desponds J, Isambert H: Preserving information while respecting privacy through an information theoretic framework for synthetic health data generation
npj Digital Medicine, 8, 49 (2025).
Pubmed
| DOI
| pdf
|
| |
|
|
|
|
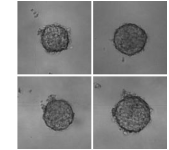
Parent C, Honari H, Tocci T, Simon F, Zaidi S, Jan A, Aubert V, Delattre O, Isambert H, Wilhelm C, Viovy J-L: Label-free Machine Learning prediction of chemotherapy on tumor spheroids using a microfluidics droplet platform
Small Science, 2500173 (2025).
Pubmed
| DOI
| pdf| supp
|
| |
|
|

|
|
Donada A, Hermange G, Tocci T, Midoun A, Prevedello G, Hadj Abed L, Dupre D, Sun W, Milo I, Tenreira Bento S, Pospori C, Innes A, Willekens C, Vargaftig J, Michonneau D, Lo Celso C, Servant N, Duffy KR, Isambert H, Cournede PH, Laplane L, Perie L: Clonal memory of cell division in humans diverges between healthy haematopoiesis and acute myeloid leukaemia
bioRxiv, 2025.06.24.660535 (2025).
Pubmed
| DOI
| bioRxiv
|
| |
|
|
|
|
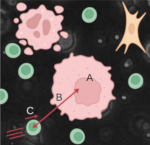
Simon F, Comes MC, Tocci T, Dupuis L, Cabeli V, Lagrange N, Mencattini A, Parrini MC, Martinelli E, Isambert H: CausalXtract: a flexible pipeline to extract causal effects from live-cell time-lapse imaging data
eLife, 13:RP95485 (2025).
Pubmed
| DOI
| bioRxiv
| pdf
| pdf+supp
[press release]
|
| |
|
|

|
|
Fusilier Z, Simon F, Calvente I, Crestey L, Clement A, Mathieu M, Jean-Marie R, Piastra-Facon F, George Clement J, Lumineau E, Tonani M, Manriquez V, Lacerda L, de Villemagne P, Piaggio E, Semetey V, Coscoy S, Martini E, Scita G, Gelly J-C, Ivaska J, Isambert H, Goudot C, Pierobon P, Lennon-Dumenil A-M, Moreau HD: Macrophages restrict tumor permissiveness to immune infiltration by controlling local collagen topography through a Tcf4-Collagen3 fibrotic axis
bioRxiv, 2025.01.17.633527 (2025).
Pubmed
| DOI
| bioRxiv
| pdf
| pdf+supp
|
| |
|
|

|
|
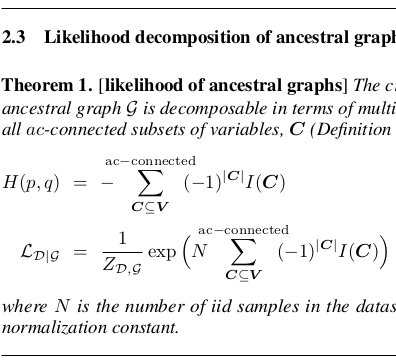
Lagrange N, Isambert H: An efficient search-and-score algorithm for ancestral graphs using multivariate information scores
opt-in article at NeurIPS 2024, arXiv, 2412.17508 (2024).
Pubmed
| DOI
| arXiv
| pdf
|
| |
|
|
|
|
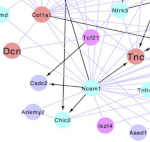
Miladinovic O, Canto P-Y, Pouget C, Piau O, Radic N, Freschu P, Megherbi A, Prats CB, Jacques S, Hirsinger E, Geeverding A, Dufour S, Petit L, Souyri M, North T, Isambert H, Traver D, Jaffredo T, Charbord P, Durand C: A multistep computational approach reveals a neuro-mesenchymal cell population in the embryonic hematopoietic stem cell niche
Development, 151, dev202614 (2024).
Pubmed
| DOI| pdf| supp
|
| |
|
|
|
|
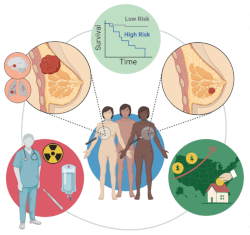
Ribeiro-Dantas MdC, Li H, Cabeli V, Dupuis L, Simon F, Hettal L, Hamy A-S and Isambert H: Learning interpretable causal networks from very large datasets, application to 400,000 medical records of breast cancer patients
iScience 27(5):109736 (2024).
Pubmed
| DOI
| arxiv
| pdf
| pdf+supp
|
| |
|
|
|
|
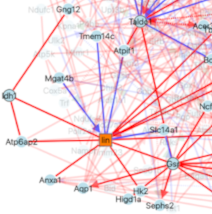
Cosgrove J, Lyne A-M, Rodriguez I, Cabeli V, Conrad C, Tenreira-Bento S, Tubeuf E, Russo E, Tabarin F, Belloucif Y, Maleki-Toyserkani S, Reed S, Monaco F, Ager A, Lobry C, Bousso P, Fernandez-Marcos PJ, Isambert H, Arguello RJ, Perie L: Metabolically primed multipotent hematopoietic progenitors fuel innate immunity
bioRxiv 2023.01.24.525166 (2023).
Pubmed
| bioRxiv
| pdf
| supp
|
| |
|
|
|
|
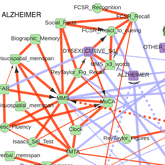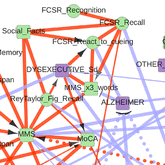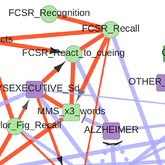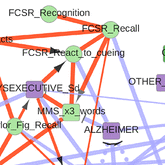
Sella N, Hamy AS, Cabeli V, Darrigues L, Lae M, Reyal F, Isambert H: Interactive exploration of a global clinical network from a large breast cancer cohort
npj Digital Medicine 5, 113 (2022).
Pubmed
| DOI
| pdf
| supp
|
| |
|
|
|
|
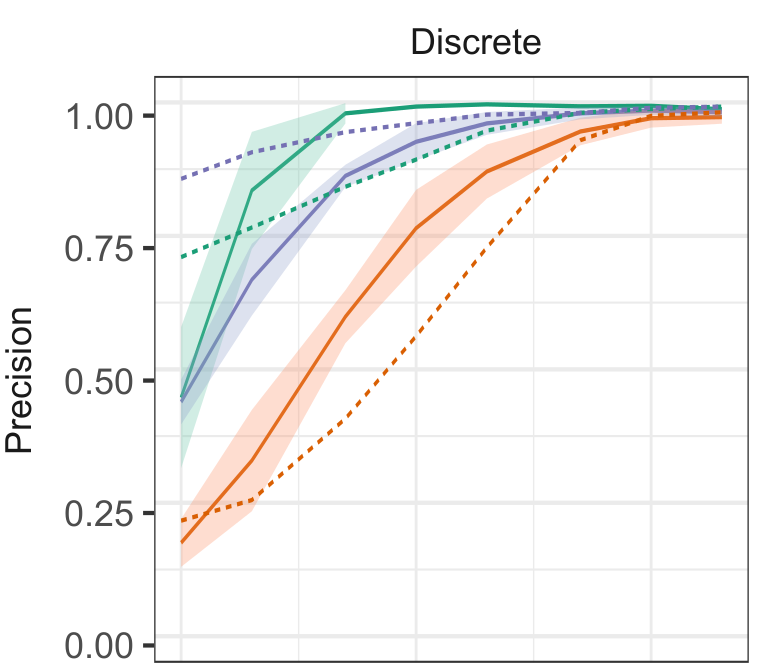
Cabeli V, Li H, Ribeiro-Dantas M, Simon F, Isambert H: Reliable causal discovery based on mutual information supremum principle for finite dataset
Why21 @ NeurIPS2021 (2021).
Pubmed
| DOI
| pdf
| supp
|
| |
|
|
|
|
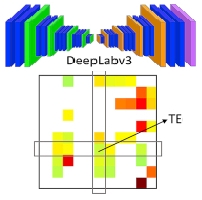
Mencattini A, Spalloni A, Casti P, Comes MC, Di Giuseppe D, Antonelli G, D Orazio M, Filippi J, Corsi F, Isambert H, Di Natale C, Longone P, Martinelli E: NeuriTES. Monitoring neurite changes through transfer entropy and semantic segmentation in bright-field time-lapse microscopy
Patterns 2(6):100261 (2021).
Pubmed
| DOI
| pdf
| supp
|
| |
|
|
|
|
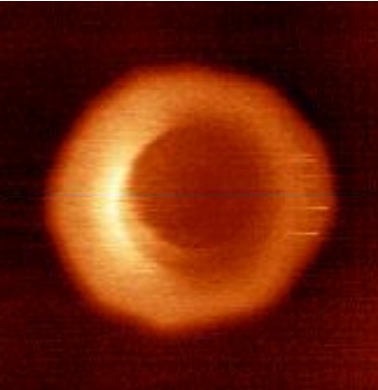
Vial A, Taveneau C, Costa L, Chauvin B, Nasrallah H, Godefroy C, Dosset P, Isambert H, Ngo KX, Mangenot S, Levy D, Bertin A, Milhiet PE: Correlative AFM and fluorescence imaging demonstrate nanoscale membrane remodeling and ring-like and tubular structure formation by septins
Nanoscale 13, 12484-12493 (2021).
Pubmed
| DOI
| pdf
| supp
|
| |
|
|
|
|
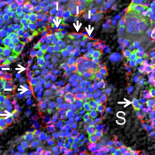
Cabeli V, Verny L, Sella N, Uguzzoni U, Verny M, Isambert H: Learning clinical networks from medical records based on information estimates in mixed-type data
PLoS Comput Biol 16(5):e1007866 (2020).
Pubmed
| DOI
| pdf
| supp
|
| |
|
|
|
|
Desterke C, Petit L, Sella N, Chevallier N, Cabeli V, Coquelin L, Durand C,
Oostendorp RAJ, Isambert H, Jaffredo T, Charbord P: Inferring gene networks in bone marrow Hematopoietic Stem Cell-supporting stromal niche populations
iScience 23(6):101222 (2020).
Pubmed
| DOI
| pdf
| supp
|
| |
|
|

|
|
Singh PP, Isambert H: Ohnologs v2: a comprehensive resource for the genes retained from whole genome duplication in vertebrates
Nucleic Acid Res 48(D1):D724-D730 (2020).
Pubmed
| DOI
| pdf
| supp
| bioRxiv
| Ohnologs server
[recommended by F1000Prime]
|
| |
|
|
|
|
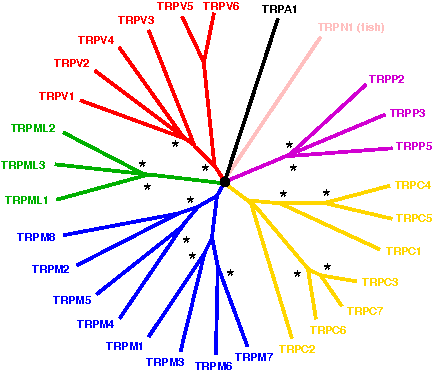
Arbabian A, Iftinca M, Altier C, Singh PP, Isambert H, Coscoy S: Mutations in calmodulin-binding domains of TRPV4/6 channels confer invasive properties to colon adenocarcinoma cells.
Channels 14(1):101-109 (2020).
Pubmed
| DOI
| pdf
| supp
|
| |
|
|

|
|
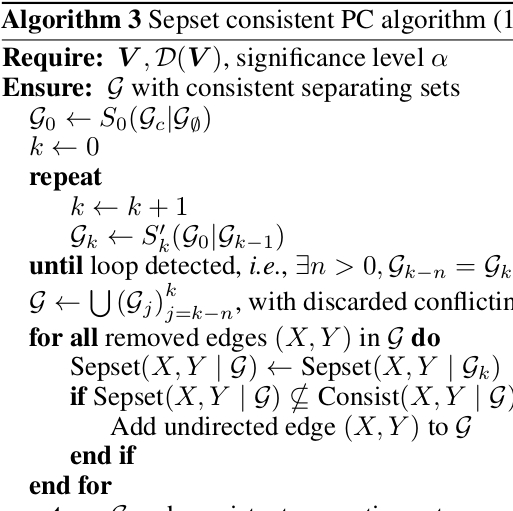
Li H, Cabeli V, Sella N, Isambert H: Constraint-based causal structure learning with consistent separating sets.
Advances in Neural Information Processing Systems (NeurIPS) 32, 14257-14266 (2019).
Pubmed
| DOI
| pdf
| supp
|
| |
|
|
|
|
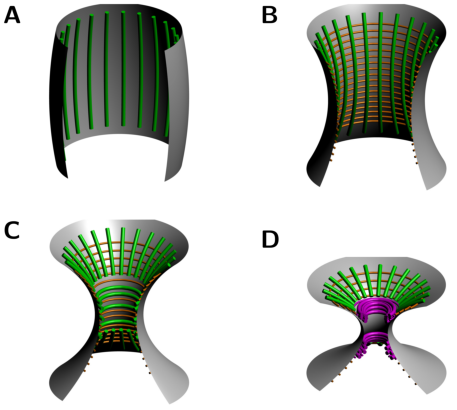
Beber A, Taveneau C, Nania M, Tsai F-C, Di Cicco A, Bassereau P, Levy D, Cabral JT, Isambert H, Mangenot S, Bertin A: Membrane reshaping by micrometric curvature sensitive septin filaments.
Nat Commun 10: 420 (2019).
Pubmed
| DOI
| pdf
| supp
|
| |
|
|
|
|
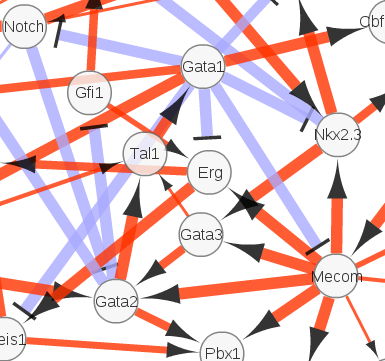
Sella N, Verny L, Uguzzoni G, Affeldt S, Isambert H: MIIC online: a web server to reconstruct causal or non-causal networks from non-perturbative data.
Bioinformatics 34(13):2311-2313 (2018).
Pubmed
| OUP
| pdf
| MIIC online server
|
| |
|
|
|
|
Verny L, Sella N, Affeldt S, Singh PP, Isambert H: Learning causal networks with latent variables from multivariate information in genomic data.
PLoS Comput Biol 13(10):e1005662 (2017).
Pubmed
| PLOS
| pdf
| supp
[recommended by F1000]
|
| |
|
|
|
|
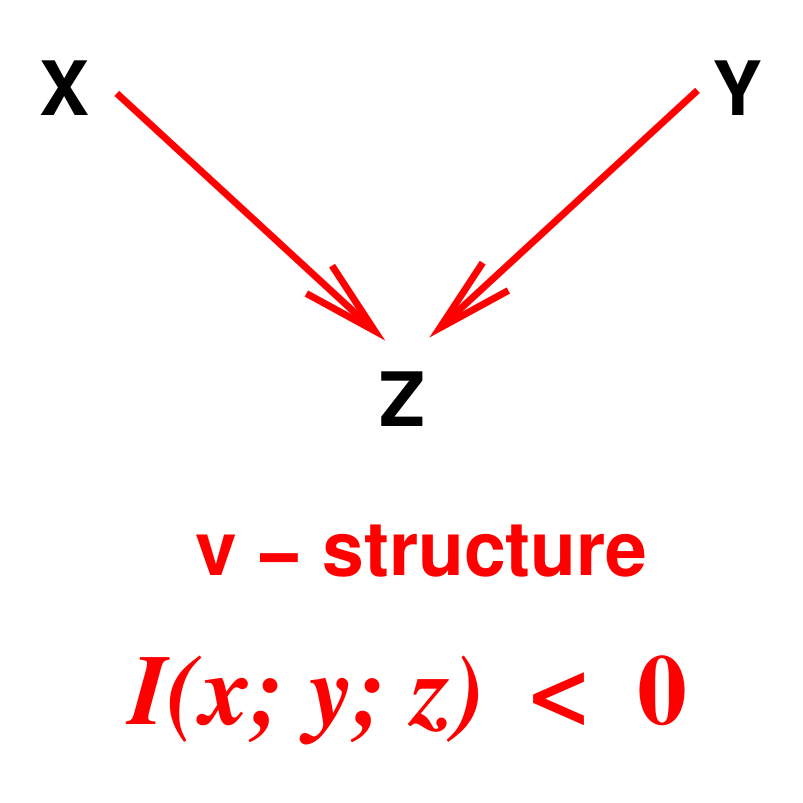
Affeldt S, Verny L, Isambert H:
3off2: A network reconstruction algorithm based on 2-point and 3-point information statistics.
BMC Bioinformatics, 17 Suppl 2:12 (2016).
Pubmed
| BMC
| pdf
| supp
|
| |
|
|
|
|
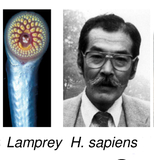
Singh P-P, Arora J, Isambert H:
Identification of ohnolog genes originating from whole genome duplication in early vertebrates, based on synteny comparison across multiple genomes.
PLoS Comput Biol, 11(7):e1004394 (2015).
Pubmed
| PLOS
| pdf
| supp
| Ohnologs server
|
| |
|
|

|
|
Affeldt S, Isambert H:
Robust reconstruction of causal graphical models based on conditional 2-point and 3-point information.
Proceedings of the 31th conference on Uncertainty in Artificial Intelligence (UAI), Amsterdam. Morgan Kaufmann (2015). [peer reviewed article]
AUAI
| pdf
| supp
|
| |
|
|
|
|
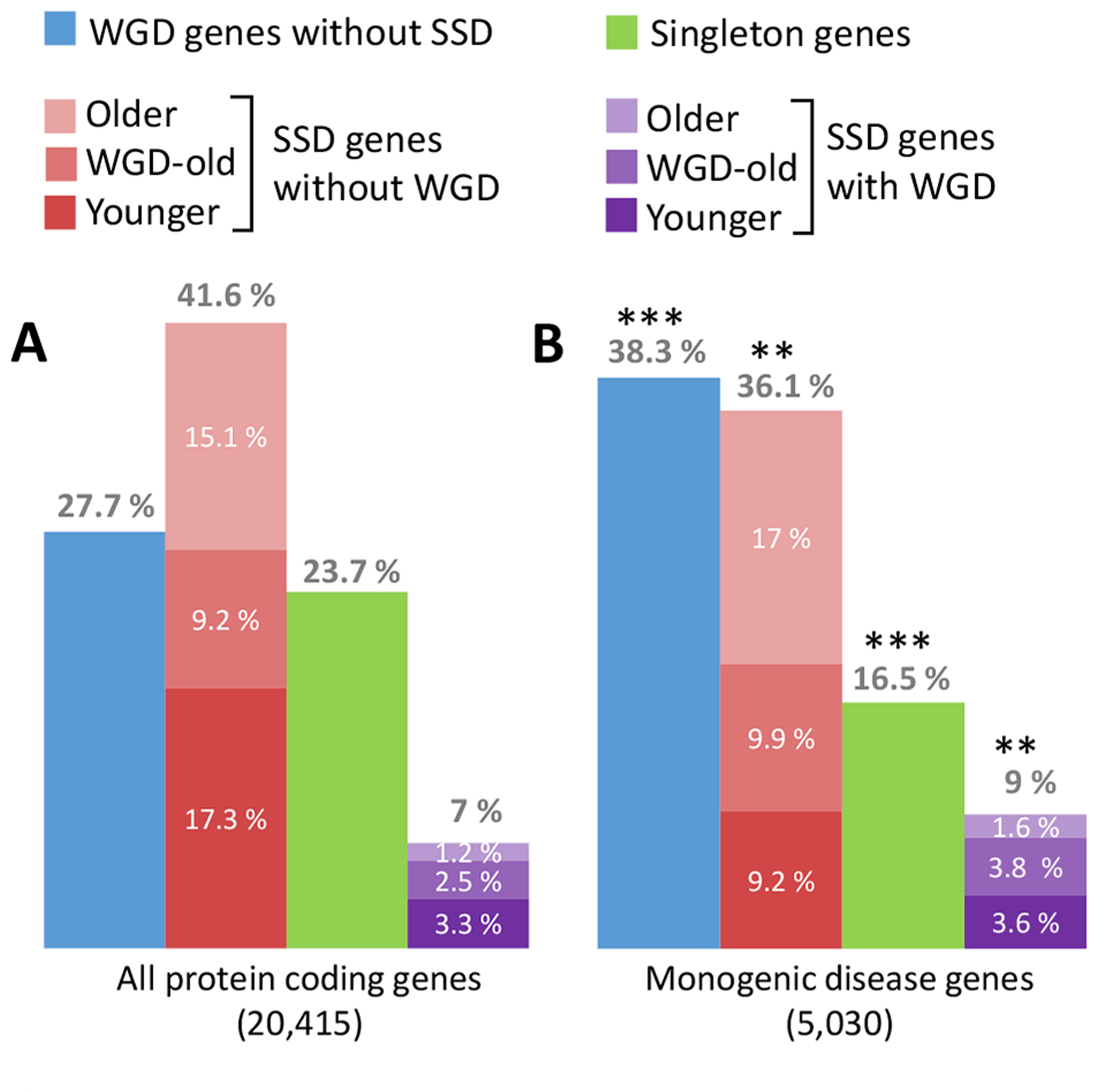
Singh PP, Affeldt S, Malaguti G, Isambert H:
Human dominant disease genes are enriched in paralogs originating from whole genome duplication.
PLoS Comput Biol, 10(7):e1003754 (2014).
Pubmed
| PLOS
| pdf
|
| |
|
|
|
|
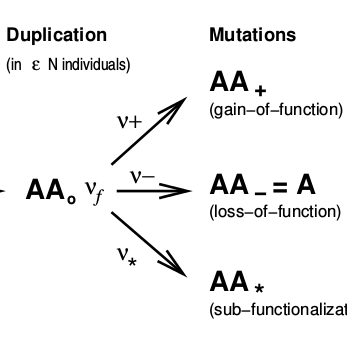
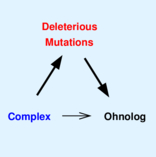
Malaguti G, Singh PP, Isambert H. On the retention of gene duplicates prone to dominant deleterious mutations.
Theor Popul Biol, 93:38-51 (2014).
Pubmed
| ScienceDirect
| pdf
|
| |
|
|
|
|
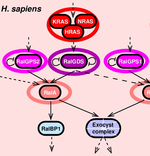
Affeldt S, Singh PP, Cascone I, Selimoglu R, Camonis J, Isambert H: Evolution and cancer: expansion of dangerous gene repertoire by whole genome duplications.
Med Sci, 29(4), 358-61 (2013) [french].
Pubmed
| EDPSciences
| pdf
|
| |
|
|
|
|
Singh PP, Affeldt S, Cascone I, Selimoglu R, Camonis J, Isambert H:
On the expansion of "dangerous" gene repertoires by whole genome duplications in early vertebrates.
Cell Rep, 2(5), 1387-1398 (2012).
Pubmed
| CellPress
| pdf
| supp
[featured by Le Point, Biofutur, CNRS]
|
| |
|
|
|
|
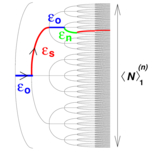
Stein RR, Isambert H:
Logistic map analysis of biomolecular network evolution.
Phys Rev E, 84, 051904 (2011).
Pubmed
| APS
| pdf
|
| |
|
|

|
|
Shukla GC, Haque F, Tor Y, Wilhelmsson LM, Toulmé JJ, Isambert H, Guo P, Rossi JJ, Tenenbaum SA, Shapiro BA:
A boost for the emerging field of RNA nanotechnology.
ACS Nano, 5(5):3405-18 (2011).
Pubmed
| ACS
| pdf
|
| |
|
|
|
|
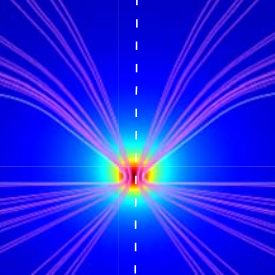
Viasnoff V, Bockelmann U, Isambert H, Laufer L, Tsori Y: Localized Joule heating produced by ion current focusing through micron-size holes.
Appl Phys Lett, 96, 163701 (2010).
AIP
| pdf
|
| |
|
|
|
|
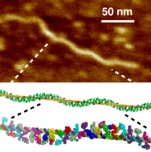
Cayrol B, Nogues C, Dawid A, Sagi I, Silberzan P, Isambert H: A nanostructure made of a bacterial noncoding RNA.
J Am Chem Soc, 131(47):17270-6 (2009).
Pubmed
| ACS
| pdf
| supp
[featured by HFSP]
|
| |
|
|
|
|
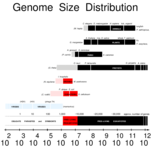
Isambert H, Stein RR:
On the need for widespread horizontal gene transfers under genome size constraint.
Biol Direct, 4:28 (2009).
Pubmed
| BMC
| pdf
|
| |
|
|
|
|
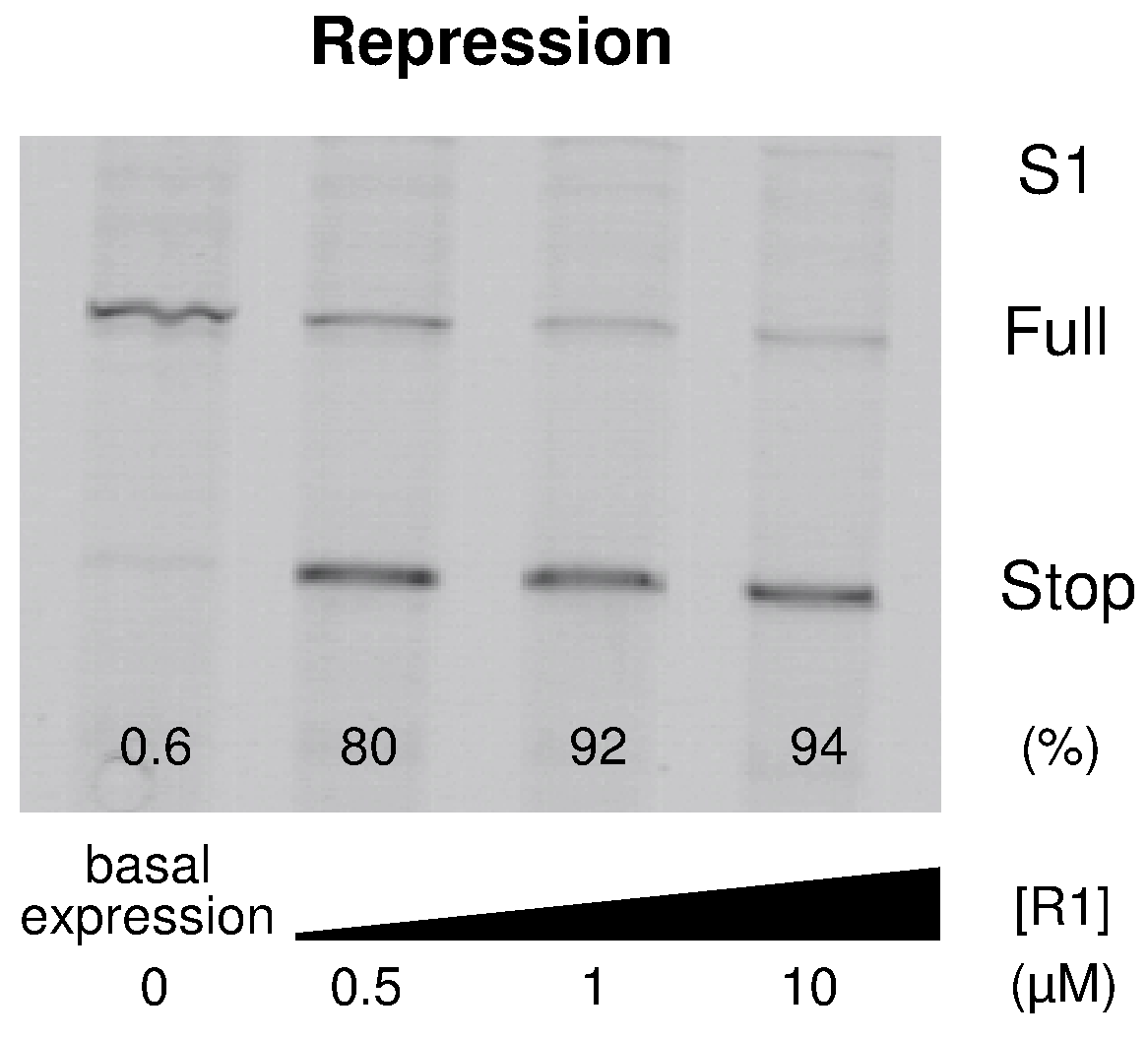
Dawid A, Cayrol B, Isambert H:
RNA synthetic biology inspired from bacteria: construction of transcription attenuators under antisense regulation.
Phys Biol, 6(2):025007 (2009).
Pubmed
| IOP
| pdf
|
| |
|
|
|
|
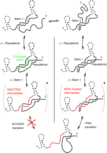
Isambert H:
The jerky and knotty dynamics of RNA.
Methods, 49(2):189-96 (2009). [review]
Pubmed
| ScienceDirect
| pdf
|
| |
|
|
|
|
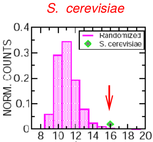
Sellerio AL, Bassetti B, Isambert H, Cosentino Lagomarsino M:
A comparative evolutionary study of transcription networks. The global role of feedback and hierachical structures.
Mol Biosyst, 5(2):170-9 (2009).
Pubmed
| RSC
| pdf
|
| |
|
|
|
|
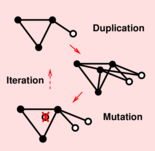
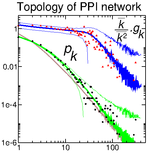
Evlampiev K, Isambert H:
Conservation and topology of protein interaction networks under duplication-divergence evolution.
Proc Natl Acad Sci USA, 105(29):9863-8 (2008).
Pubmed
| PNAS
| pdf
| supp
|
| |
|
|
|
|
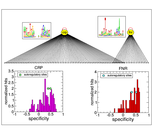
Evlampiev K, Isambert H:
Modeling protein network evolution under genome duplication and domain shuffling.
BMC Syst Biol, 1:49 (2007).
Pubmed
| BMC
| pdf
| supp
|
| |
|
|
|
|
Cosentino Lagomarsino M, Jona P, Bassetti B, Isambert H:
Hierarchy and feedback in the evolution of the Escherichia coli transcription network.
Proc Natl Acad Sci USA, 104(13):5516-20 (2007).
Pubmed
| PNAS
| pdf
| supp
[featured by F1000]
|
| |
|
|
|
|
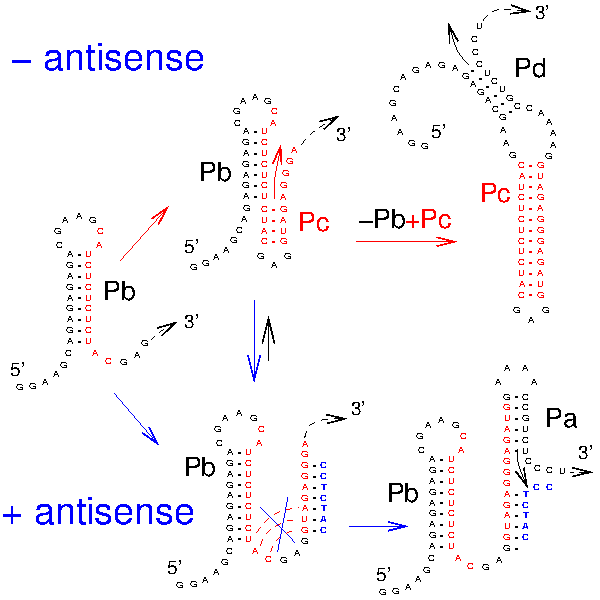
Xayaphoummine A, Viasnoff V, Harlepp S, Isambert H:
Encoding folding paths of RNA switches.
Nucleic Acids Res, 35(2):614-22 (2007).
Pubmed
| OXFORD
| pdf
[featured by HFSP]
|
| |
|
|
|
|
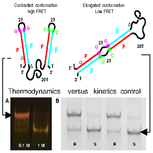
Viasnoff V, Meller A, Isambert H:
DNA nanomechanical switches under folding kinetics control.
Nano Lett, 6(1):101-4 (2006).
Pubmed
| ACS
| pdf
[featured by Nanowerk]
|
| |
|
|
|
|
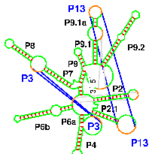
Xayaphoummine A, Bucher T, Isambert H:
Kinefold web server for RNA/DNA folding path and structure prediction including pseudoknots and knots.
Nucleic Acids Res, 33:W605-10 (2005).
Pubmed
| OXFORD
| pdf
| supp
| Kinefold server
|
| |
|
|
|
|
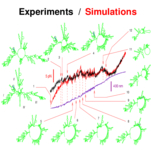
Harlepp S, Marchal T, Robert J, Léger JF, Xayaphoummine A, Isambert H, Chatenay D:
Probing complex RNA structures by mechanical force.
Eur Phys J E, 12(4):605-15 (2003).
Pubmed
| Springer
| pdf
|
| |
|
|
|
|
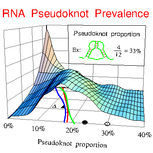
Xayaphoummine A, Bucher T, Thalmann F, Isambert H:
Prediction and statistics of pseudoknots in RNA structures using exactly clustered stochastic simulations.
Proc Natl Acad Sci USA, 100(26):15310-5 (2003).
Pubmed
| PNAS
| pdf
|
| |
|
|
|
|
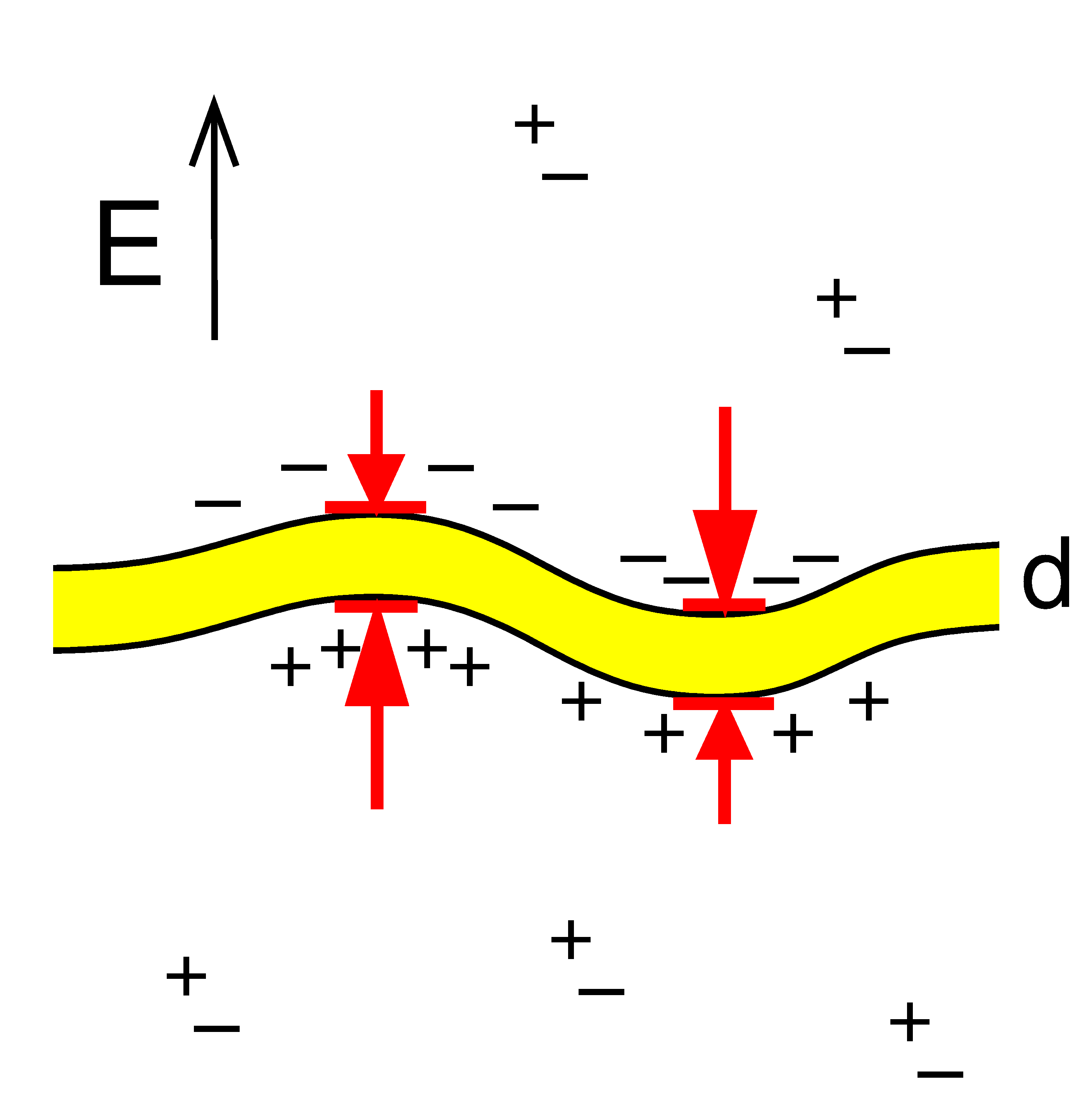
Sens P, Isambert H:
Undulation instability of lipid membranes under an electric field.
Phys Rev Lett, 88(12):128102 (2002).
Pubmed
| APS
| pdf
[featured by NaturePhysics]
|
| |
|
|

|
|
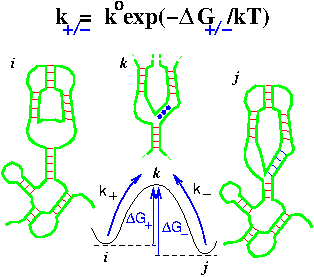
Isambert H:
Voltage addressable nanomemories in DNA?
C R Physique, 3, 391-396 (2002).
ScienceDirect
| pdf
|
| |
|
|
|
|
Isambert H, Siggia ED:
Modeling RNA folding paths with pseudoknots: application to hepatitis delta virus ribozyme.
Proc Natl Acad Sci USA, 97(12):6515-20 (2000).
Pubmed
| PNAS
| pdf
[featured by Science]
|
| |
|
|

|
|
Magnúsdóttir S, Isambert H, Heller C, Viovy JL:
Electrohydrodynamically induced aggregation during constant and pulsed field capillary electrophoresis of DNA.
Biopolymers, 49(5):385-401 (1999).
Pubmed
| Wiley
| pdf
|
| |
|
|
|
|
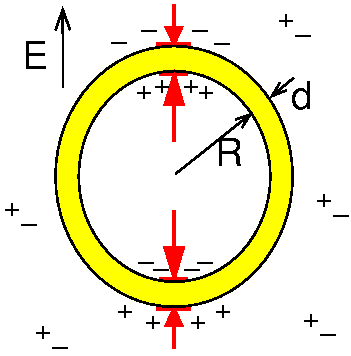
Isambert H:
Understanding the electroporation of cells and artificial bilayer
membranes.
Phys Rev Lett, 80 (15), 3404 (1998).
APS
| pdf
|
| |
|
|

|
|
Ladoux B, Isambert H, Léger JF, Viovy JL:
A new rectifying electrohydrodynamic instability at the boundary of charged gels.
Phys Rev Lett, 81, 3793 (1998).
APS
| pdf
|
| |
|
|
|
|
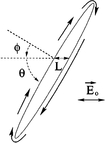
Isambert H, Ajdari A, Viovy JL, Prost J:
Electrohydrodynamic patterns in macroion dispersions under a strong electric field.
Phys Rev E, 56, 5688 (1997).
APS
| pdf
|
| |
|
|
|
|
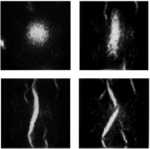
Isambert H, Ajdari A, Viovy JL, Prost J:
Electrohydrodynamic patterns in charged colloidal solutions.
Phys Rev Lett, 78 (5), 971 (1997).
APS
| pdf
|
| |
|
|
|
|
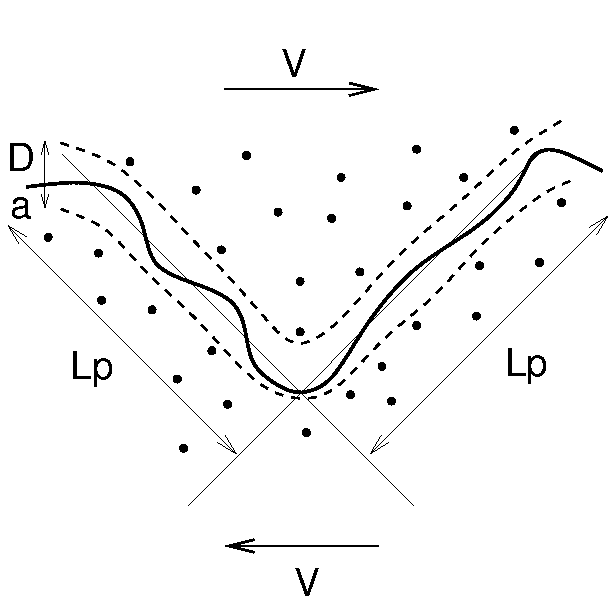
Isambert H, Maggs AC:
Dynamics and rheology of actin solutions.
Macromolecules, 29(3), 1036 (1996).
ACS
| pdf
|
| |
|
|

|
|
Isambert H, Maggs AC:
Bending of actin filaments.
Europhys Lett, 31, 263 (1995).
IOP
| pdf
|
| |
|
|
|
|
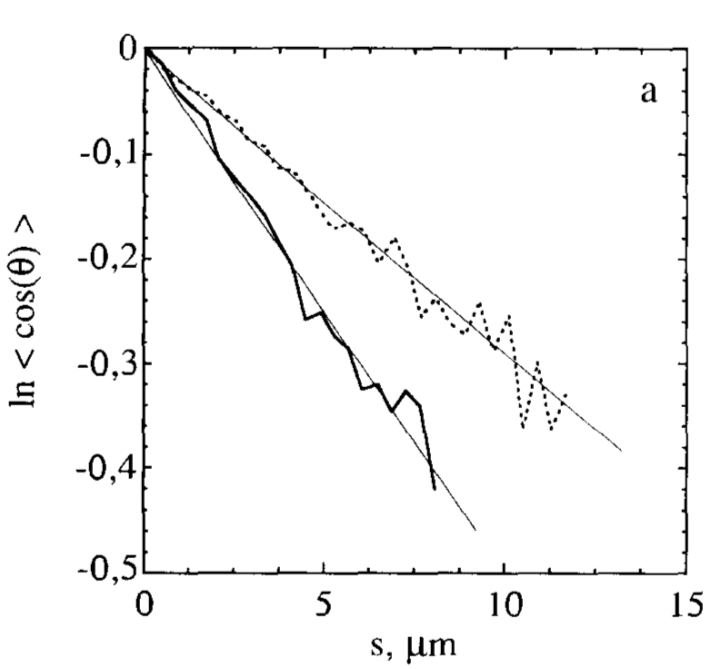
Isambert H, Venier P, Maggs AC, Fattoum A, Kassab R, Pantaloni D, Carlier MF:
Flexibility of actin filaments derived from thermal fluctuations.
J Biol Chem, 270(19):11437-44 (1995).
Pubmed
| ASBMB
| pdf
|





























































